DNA Conjugation Services
Sequence-Defined DNA FunctionalizationFlexible Coupling for Probes, Barcodes, and Hybrid ReagentsFor Assays, Imaging, and Nanobiotechnology
Build DNA constructs that do more than hybridize. Our DNA conjugation services support custom covalent attachment of fluorophores, quenchers, biotin, peptides, proteins, antibodies, small molecules, polymers, and particle surfaces to single-stranded or double-stranded DNA formats used in assay development, biosensing, spatial biology, and nanobiotechnology. We work from sequence and application requirements to choose the right handle, linker, and purification route for research-grade conjugates that are easier to interpret and easier to reproduce.
Projects can be configured around 5', 3', or internal modification sites, with chemistries selected according to payload class, buffer compatibility, steric constraints, and downstream readout. Whether you need a simple labeled probe or a more complex antibody–DNA, protein–DNA, bead–DNA, or small-molecule–DNA construct, we combine conjugation design, purification strategy, and analytical characterization in one workflow. For adjacent needs, our broader oligonucleotide bioconjugation and DNA labeling services can also be integrated into the same project path.
How DNA Conjugation Solves Real Experimental Bottlenecks
DNA conjugation turns a sequence-defined oligonucleotide into a functional tool matched to the way your assay or model actually works. By attaching the right label, affinity tag, biomolecule, or surface handle at the right position, DNA can shift from a simple hybridization element to a capture reagent, barcode carrier, signal amplifier, targeting handle, or programmable assembly component. This is especially important when projects depend on low background, preserved hybridization, controlled stoichiometry, and reliable downstream readout rather than merely obtaining a modified strand on paper.
In practice, well-designed DNA conjugation helps teams avoid the common trade-off between functionality and usability. A fluorophore must remain bright without disrupting duplex formation; a protein or antibody partner must retain activity after coupling; a bead- or nanoparticle-bound oligo must stay accessible for hybridization; and a multiplex barcode panel must be consistent enough to compare across experiments and lots. The value of the service lies in making those constraints visible early and building the conjugate around them.
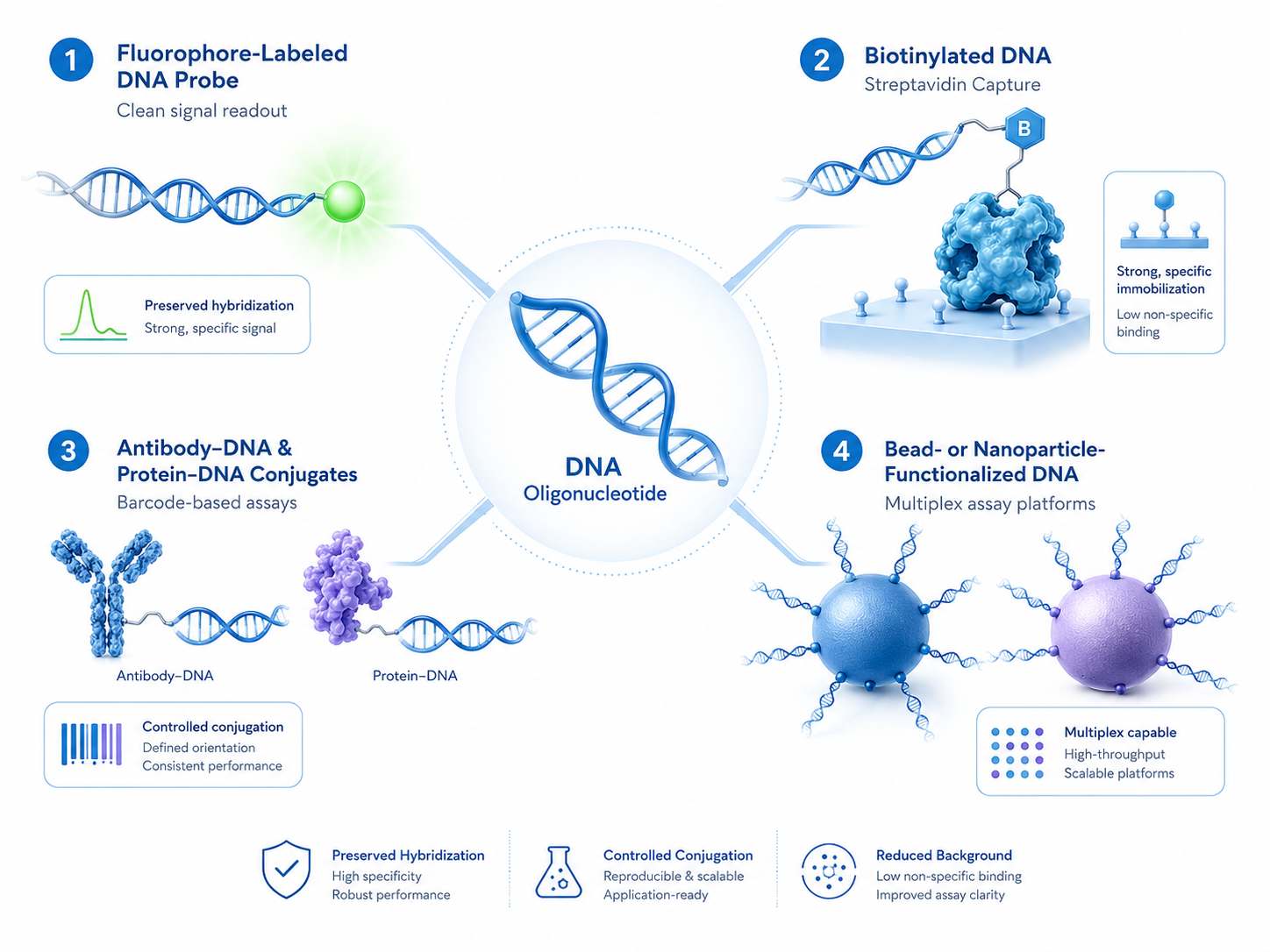 Schematic view of DNA conjugation strategies that improve signal generation, target capture, and assay compatibility across labeled probes, barcoded affinity reagents, and surface-bound DNA constructs.
Schematic view of DNA conjugation strategies that improve signal generation, target capture, and assay compatibility across labeled probes, barcoded affinity reagents, and surface-bound DNA constructs.Common Challenges in DNA Conjugation Projects
A DNA sequence may be available with an amino, thiol, azide, or alkyne handle, but that does not automatically make it compatible with the partner molecule you want to attach. We help align the reactive group, conjugation sequence, and buffer conditions with the real chemical behavior of dyes, proteins, peptides, and surfaces so that coupling efficiency is not lost at the first step.
DNA conjugates often fail when the attachment site introduces steric crowding, excess hydrophobicity, or a linker that is too short or too labile. Our design workflows focus on preserving duplex formation, target accessibility, fluorophore performance, and biomolecule activity while still achieving useful conjugation levels.
In barcode-driven and signal-amplified assays, residual free dye, free DNA, or unconjugated protein can distort readout and reduce reproducibility. We plan purification and analytical controls around separation of the desired conjugate from excess small molecules, unconjugated oligo, and unconjugated biomolecule rather than treating cleanup as an afterthought.
A conjugate that works once at screening scale is not necessarily suitable for a barcode panel, validation series, or repeat order. We emphasize route selection, batch comparability, and fit-for-purpose QC so that DNA conjugates remain interpretable when projects move from one-off preparation to broader assay deployment.
Our DNA Conjugation Service Modules
We support custom DNA conjugation projects ranging from simple labeled oligos to multi-component hybrid constructs. Service scope can include design review, reactive handle selection, DNA synthesis or customer-supplied starting materials, coupling, purification, analytical characterization, and application-oriented documentation.
 Fluorophore, Quencher, and Reporter DNA Conjugation
Fluorophore, Quencher, and Reporter DNA Conjugation
Capabilities include:
- 5', 3', or internal labeling of DNA with fluorescent dyes, dark quenchers, and other reporter groups
- Design support for hybridization probes, molecular beacons, FRET pairs, and imaging barcodes
- Spacer and linker selection to reduce dye-induced interference with duplex formation
- Purification options selected according to sequence, modification load, and downstream sensitivity
- UV-Vis and chromatographic assessment of labeling quality
Typical applications:
qPCR and probe development, hybridization assays, fluorescence readouts, and custom barcode oligos
 Biotinylated and Affinity-Tagged DNA
Biotinylated and Affinity-Tagged DNA
Capabilities include:
- Terminal or internal biotin installation for capture, pull-down, immobilization, or assay assembly
- Design of spacer lengths to improve streptavidin accessibility and reduce steric masking
- Support for capture oligos, bait strands, and affinity-enabled assay controls
- Purity control for applications sensitive to free biotin or incomplete modification
- Compatibility planning for downstream surfaces, beads, and multi-step workflows
Typical applications:
Target capture, enrichment workflows, surface immobilization, and affinity assay development
 Antibody–DNA and Protein–DNA Conjugates
Antibody–DNA and Protein–DNA Conjugates
Capabilities include:
- DNA coupling to antibodies, enzymes, and other proteins using fit-for-purpose amine, thiol, or click-based strategies
- Route selection to balance conjugation efficiency with preservation of binding or enzymatic function
- Conjugate loading control and purification planning to reduce free barcode and unconjugated partner
- Integration with related protein–DNA conjugation needs for custom protein formats
- Analytical packages tailored for barcode assays, proximity methods, and affinity reagent development
Typical applications:
Multiplex imaging, proximity assays, immuno-PCR, biosensing, and DNA-barcoded reagent development
 Peptide, Small-Molecule, and Polymer-Linked DNA
Peptide, Small-Molecule, and Polymer-Linked DNA
Capabilities include:
- DNA conjugation to peptides, small molecules, polymers, and related synthetic partners
- Route planning for hydrophobic or sterically demanding payloads that can affect oligo behavior
- Optional linkage strategies for improved spacing, solubility, or downstream compatibility
- Access to related peptide–oligonucleotide conjugation and PEG-conjugated oligonucleotide workflows
- Characterization focused on conjugation integrity and usable recovery
Typical applications:
Delivery research tools, ligand-enabled DNA constructs, programmable assemblies, and custom hybrid reagents
 DNA Functionalization for Beads, Surfaces, and Nanoparticles
DNA Functionalization for Beads, Surfaces, and Nanoparticles
Capabilities include:
- DNA attachment to gold nanoparticles, bead systems, and assay-compatible surfaces
- Support for barcode display, capture orientation, and hybridization accessibility considerations
- Project fit with Luminex bead labeled DNA and other particle-based assay formats
- Purification and washing strategies selected for surface-bound or colloidal systems
- Characterization planning for lot-to-lot consistency and functional attachment
Typical applications:
Bead assays, microarray-style capture systems, nanobiosensors, and programmable DNA-decorated materials
 Conjugation Design, Purification, and QC Support
Conjugation Design, Purification, and QC Support
Capabilities include:
- Feasibility review for sequence, modification position, partner molecule, and intended use
- Selection of coupling order, orthogonal handles, and purification workflow
- HPLC, PAGE, LC-MS, UV-Vis, and other fit-for-purpose analytical options
- Assessment of free label removal, conjugation integrity, and loading uniformity
- Data packages that help research teams compare batches and progress follow-on work
Deliverables:
Conjugation summary, chromatographic or spectrometric data, purity assessment, and project-specific technical reporting
Key Design Parameters for Custom DNA Conjugates
Successful DNA conjugation depends less on a single reaction name and more on how sequence, attachment site, payload, and purification strategy work together. The table below summarizes the design variables that most often determine whether a conjugate is easy to make, easy to purify, and useful in the intended assay.
| Design Parameter | Common Options | What to Evaluate | Impact on Final Conjugate | Why It Matters to the Project |
| DNA Format | ssDNA oligo, dsDNA fragment, barcode strand, aptamer-like construct | Sequence length, secondary structure, and hybridization role in the assay | Influences synthesis route, purification choice, and conjugation accessibility | Determines how much structural flexibility is available for modification |
| Attachment Position | 5' end, 3' end, or internal modification site | Distance from hybridizing region, protein-binding motif, or cleavage-sensitive region | Affects steric crowding, duplex formation, and signal behavior | Often the main factor behind "conjugated but not functional" outcomes |
| Reactive Handle | Amino, thiol, azide, alkyne, DBCO-ready, or other orthogonal handle | Compatibility with payload chemistry, solvent limits, and buffer composition | Controls coupling efficiency, selectivity, and side-product profile | Prevents avoidable rework caused by handle–payload mismatch |
| Linker or Spacer | Short alkyl spacer, PEG-type spacer, rigid or flexible linker | Whether the payload needs distance from the DNA backbone to remain accessible | Influences solubility, steric exposure, and readout quality | Critical when coupling bulky dyes, proteins, or surfaces |
| Payload Class | Fluorophore, biotin, peptide, protein, antibody, small molecule, bead, or nanoparticle | Size, charge, hydrophobicity, and sensitivity to reaction conditions | Drives route selection, purification difficulty, and analytical method fit | Different payloads create very different technical risks even on the same DNA sequence |
| Purification Target | Desalting, HPLC, PAGE, SEC, or combined purification workflow | Need to remove free label, truncated oligos, or unconjugated biomolecule | Determines background level and lot-to-lot reproducibility | Especially important for multiplex panels and high-sensitivity assays |
Common DNA Conjugation Strategies and When They Fit Best
There is no universal conjugation method for every DNA project. The most practical route depends on whether the project needs speed, site preference, biomolecule compatibility, or tighter control over heterogeneous species. The table below summarizes the approaches most commonly evaluated for custom DNA conjugation workflows.
| Conjugation Strategy | Technical Approach | Best Suited For | Key Considerations |
| On-Synthesis Modification Incorporation | Label or reactive handle introduced during DNA synthesis using modified phosphoramidites or solid supports | Standardized terminal or internal labels and handles | Strong positional control; often the most direct route when the required modification is synthesis-compatible |
| Amine-Reactive Coupling | Amino-modified DNA reacted with activated esters or related amine-reactive payloads | Dyes, small molecules, and selected linker installations | Practical and widely used, but requires attention to hydrolysis, buffer components, and excess reagent removal |
| Thiol–Maleimide Coupling | Thiol-modified DNA coupled to maleimide-bearing partners under controlled conditions | Protein, peptide, and selected surface-linker projects | Useful for targeted sulfhydryl chemistry, but thiol handling, reduction state, and side reactions must be managed |
| CuAAC Click Chemistry | Azide–alkyne cycloaddition used for efficient post-synthetic conjugation | Robust small-molecule labeling and modular DNA conjugate assembly | High efficiency and modularity; copper compatibility with the payload must be considered |
| SPAAC / Copper-Free Click Chemistry | Strain-promoted cycloaddition between azide and cyclooctyne-type handles such as DBCO | Protein, antibody, and other copper-sensitive biomolecule conjugates | Mild and bioorthogonal; valuable when preserving biomolecule performance is more important than reagent economy |
| Surface and Particle Functionalization | DNA coupled or immobilized onto activated beads, nanoparticles, or assay surfaces using application-specific chemistry | Capture probes, bead assays, microarray-style systems, and nanobiosensors | Surface density, accessibility, and wash stability matter as much as the coupling reaction itself |
Analytical Characterization Framework for DNA Conjugates
DNA conjugation projects benefit from analytical plans tailored to the construct type rather than a one-size-fits-all release test. A dye-labeled oligo, an antibody–DNA barcode reagent, and a bead-bound capture strand do not fail in the same way. Our QC framework therefore combines core identity testing with application-relevant checks for purity, free-species removal, loading, and functional compatibility.
| Analytical Category | Methodology | Purpose in Project Evaluation | Typical Data Delivered |
| Molecular Identity | LC-MS, MALDI-MS, or other mass-based confirmation as appropriate to construct type | Confirms that the intended DNA modification or conjugate mass is present | Mass spectra, deconvoluted mass values, identity summary |
| Purity and Free-Species Assessment | Analytical HPLC, ion-exchange methods, PAGE, or capillary-based separation | Evaluates removal of unconjugated DNA, free label, and related side products | Chromatograms or electropherograms, purity profile, impurity observations |
| Loading or Labeling Evaluation | UV-Vis analysis, ratio assessment, gel-shift comparison, or construct-specific calculation | Estimates whether the DNA or payload is attached at the expected level | Absorbance data, loading estimate, comparative summary |
| Macromolecular Heterogeneity Check | SEC, SDS-PAGE, gel-shift, or particle-relevant characterization when needed | Reviews conjugate distribution for protein-, antibody-, and particle-linked DNA formats | Distribution profile, band pattern, or size-related data package |
| Hybridization and Functional Compatibility | Duplex formation check, capture test, binding assay, or assay-specific feasibility evaluation | Confirms the conjugate still behaves as required in the intended workflow | Functional readout summary and project-specific interpretation |
| Stability-Oriented Assessment | Defined-condition monitoring of selected samples over time | Identifies obvious degradation, dissociation, or signal drift risks | Time-point comparison and storage guidance notes |
| Lot Comparison | Side-by-side analytical review across repeat preparations | Supports panel build, repeat orders, and cross-batch reproducibility decisions | Overlay plots, comparative metrics, lot summary |
| Reporting Package | Consolidated technical summary with method-specific outputs | Helps internal teams review construct quality and next-step readiness | Technical report, key raw data excerpts, and interpretation notes |
DNA Conjugation Workflow from Design Review to Deliverable
We begin by reviewing DNA format, sequence constraints, intended application, partner molecule, preferred scale, and any existing modification handles. This step identifies early risks such as incompatible buffer systems, steric issues, and purification challenges before materials are committed.
Conjugation position and chemistry are selected according to how the construct needs to function after coupling. We define whether 5', 3', or internal modification is most appropriate, which reactive handle best matches the payload, and whether a spacer is needed to preserve accessibility or signal quality.
DNA and the conjugation partner are prepared in a route-compatible form, whether from custom synthesis, customer-supplied material, or a partially modified intermediate. Reaction conditions are planned to minimize unnecessary exposure of sensitive partners to harsh solvents, metals, or reducing conditions.
The conjugation reaction is performed under controlled conditions with attention to stoichiometry, order of addition, and handle integrity. In-process checks help determine whether the route is progressing toward a usable conjugate or needs adjustment before purification.
Cleanup is selected around the impurity profile of the project, not by habit. Depending on the construct, this may mean removing excess small-molecule reagent, separating unconjugated DNA from conjugated species, or resolving free biomolecule from the desired DNA-linked fraction.
Final deliverables can include analytical confirmation, purity assessment, loading-related data, and project notes relevant to the intended assay. When needed, we also support repeat preparation, panel expansion, or route refinement for the next phase of the program.
Why Teams Choose Our DNA Conjugation Service
We evaluate sequence behavior, attachment position, and downstream hybridization before selecting chemistry, which helps prevent designs that look feasible synthetically but fail functionally.

Our workflows can be adapted for dyes, affinity tags, proteins, peptides, polymers, beads, and nanoparticles, allowing the conjugation route to follow the project need rather than forcing everything through one platform.
We pay close attention to free label, unconjugated DNA, and unconjugated biomolecule because those species often create more assay problems than the coupling reaction itself.
Keeping route design, coupling, purification, and analytics in one service path reduces handoff friction and gives project teams a clearer basis for troubleshooting, repeat orders, and panel growth.
Application Areas for Custom DNA Conjugates
Fluorescent Probes and Hybridization Assays
- Labeled DNA probes for sequence recognition and signal generation.
- FRET-ready or quencher-enabled constructs for assay optimization.
- Probe architectures designed to preserve hybridization performance.
Multiplex Imaging and DNA-Barcoded Affinity Reagents
- Antibody–DNA and protein–DNA conjugates for barcode-driven detection workflows.
- Panel-oriented preparations where free barcode control matters.
- Constructs designed for high-content readout and iterative imaging schemes.
Immuno-PCR and Proximity-Based Detection
- DNA-linked biomolecules used to convert target recognition into amplifiable signal.
- Conjugates that balance binding preservation with low-background barcode performance.
- Project support for reagent build, cleanup, and analytical review.
Capture, Pull-Down, and Surface Immobilization
- Biotinylated or otherwise affinity-enabled DNA for target capture workflows.
- Surface-bound DNA formats for microarray-style or plate-based systems.
- Designs that improve accessibility after immobilization.
DNA Nanotechnology and Programmable Assemblies
- DNA linked to peptides, proteins, or small molecules for controlled assembly.
- Hybrid constructs used in nanobiosensing and molecular organization studies.
- Conjugation routes selected to preserve programmability of the DNA scaffold.
Bead and Nanoparticle Functionalization
- DNA-decorated particles for capture, barcoding, or signal amplification.
- Bead-linked oligos for multiplex assay workflows.
- Attachment strategies chosen for stable display and usable hybridization access.
Plan Your DNA Conjugation Project with a Service Team That Understands the Chemistry
Whether you are developing a labeled DNA probe, building a DNA-barcoded affinity reagent, or functionalizing a surface with custom oligos, we can help you define the right route before avoidable chemistry decisions slow the project down.
We support design review, coupling strategy selection, purification planning, and analytical characterization for DNA conjugates that need to work in real experimental workflows. For narrower project types, you can also explore our antibody–DNA conjugation, protein–DNA conjugation, biotin-labeled DNA service, and fluorescence labeling of oligonucleotides pages.Contact our scientific team to discuss your DNA conjugation requirements, preferred modification strategy, and desired analytical package.
Frequently Asked Questions (FAQ)
Single-stranded DNA oligos, double-stranded DNA fragments, barcode strands, capture probes, and other custom DNA constructs can all be considered, provided the sequence, handle position, and downstream application are defined.
Yes. Site selection depends on whether the modification must stay away from the hybridizing region, remain exposed after immobilization, or avoid interfering with a protein- or ligand-binding motif.
That depends on the available functional groups and how sensitive the biomolecule is to reaction conditions. Amine-reactive, thiol–maleimide, and copper-free click workflows are common starting points, but the best choice is application-specific.
Purification is chosen around the impurity profile of the project. Depending on the construct, this may involve HPLC, PAGE, SEC, desalting, or combined workflows to separate the desired conjugate from excess reagents and unconjugated species.
Yes. DNA conjugation is frequently used to build barcoded affinity reagents and related panel components, with particular attention to barcode purity, conjugate loading, and batch comparability.

