Peptide Oligonucleotide Conjugation
Targeted Intracellular DeliverySite-Specific Peptide–Oligo AssemblyCustom Support for ASO, siRNA, PMO, DNA & RNA Programs
Accelerate complex nucleic acid projects with custom peptide oligonucleotide conjugation designed for research, discovery, and preclinical development teams. Peptide–oligonucleotide conjugates (POCs) combine the sequence-selective function of oligonucleotides with the transport, targeting, or membrane-interacting properties of peptides, making them valuable when unconjugated oligos show limited uptake, insufficient intracellular trafficking, or poor compatibility with the intended biological model.
We support peptide conjugation across a broad range of oligonucleotide classes, including ASOs, siRNA, splice-switching oligonucleotides, PMO, DNA, RNA, and selected probe formats. Projects are tailored around peptide class, oligonucleotide chemistry, attachment site, linker strategy, and analytical requirements so that the final construct is not only synthetically accessible, but also aligned with the desired mechanism, assay path, and downstream development plan. For broader oligonucleotide bioconjugation needs, the same workflow can be extended to other ligand and payload combinations.

What Problems Can Peptide Oligonucleotide Conjugation Solve?
Many oligonucleotide programs fail to translate cleanly from sequence design to functional delivery because the payload reaches the wrong compartment, enters cells inefficiently, or loses activity after an otherwise successful conjugation step. Peptide oligonucleotide conjugation is often explored when teams need a more defined molecular format than nanoparticle complexes, but still require better cellular entry, receptor engagement, tissue preference, endosomal processing, or assay-compatible tracking. Depending on the project, the peptide component may serve as a cell-penetrating motif, receptor-targeting ligand, endosomolytic helper, nuclear localization element, or affinity/handle sequence for capture and assembly workflows. The real challenge is not simply linking two molecules together; it is building a conjugate that preserves oligonucleotide function, controls stoichiometry, remains purifiable, and produces data that can support confident go/no-go decisions.
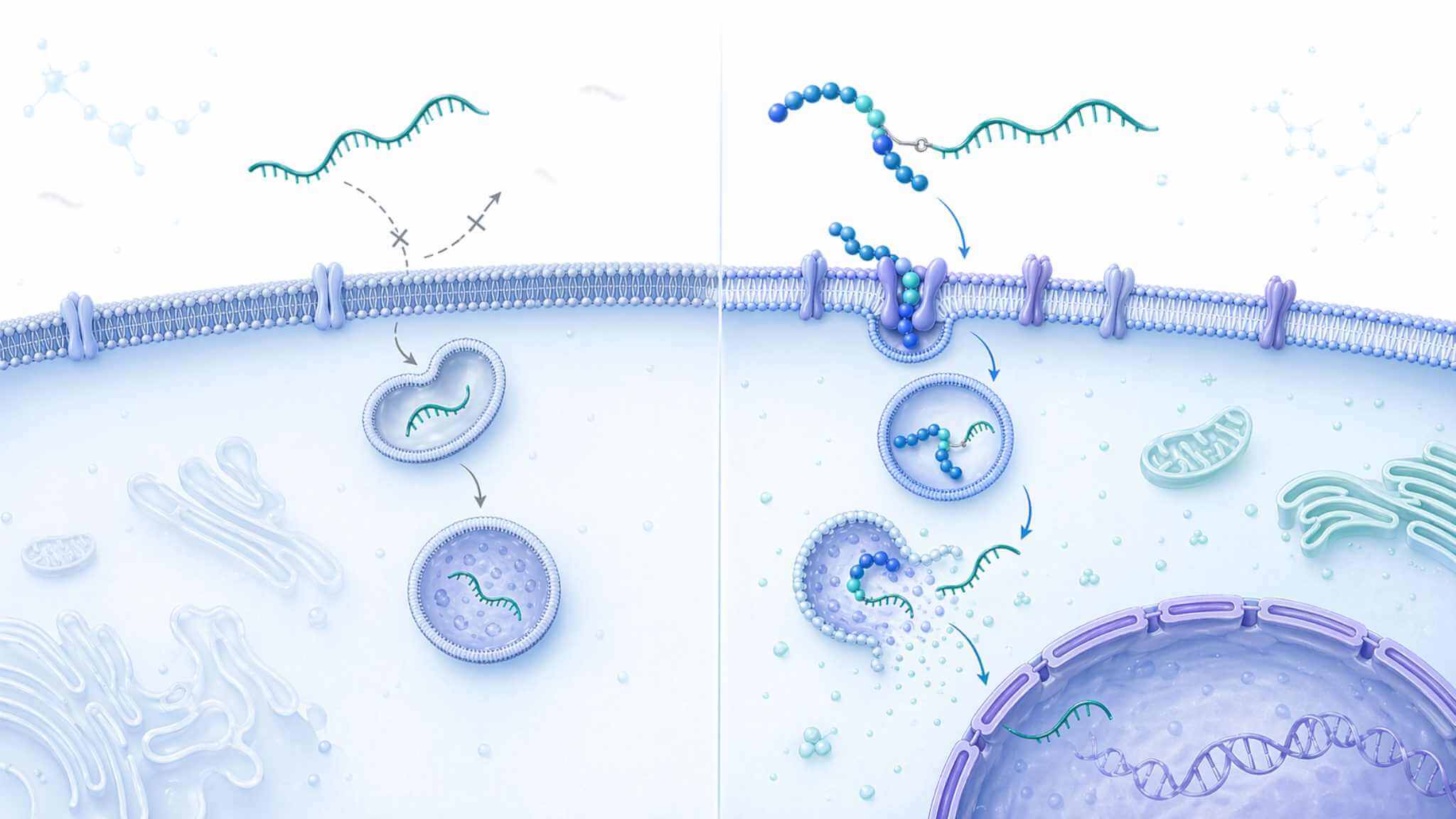 Conceptual schematic showing how a peptide–oligonucleotide conjugate can improve uptake and intracellular routing while maintaining a defined, analyzable conjugate format.
Conceptual schematic showing how a peptide–oligonucleotide conjugate can improve uptake and intracellular routing while maintaining a defined, analyzable conjugate format.Key Technical Challenges Clients Commonly Need to Resolve
A cationic CPP, receptor-targeting peptide, endosomal escape motif, or localization sequence can produce very different uptake and trafficking behavior. We help map peptide choice to the intended biology instead of treating all peptide additions as interchangeable.
Attachment at the wrong terminus, excessive steric bulk, or an unstable linker can reduce hybridization performance, interfere with RNAi loading, or compromise steric-blocking function. Conjugation design is therefore evaluated together with oligo architecture rather than as a separate chemistry step.
Projects often require a defined 1:1 construct rather than a mixed population of partially modified species. We use handle planning and site-selective coupling strategies to reduce ambiguity in attachment position and conjugation ratio.
Arginine-rich or hydrophobic peptides can change chromatographic behavior, increase self-association, and complicate formulation or assay performance. Early assessment of sequence composition and linker spacing helps avoid late-stage purification surprises.
Unconjugated oligo, truncated peptide, hydrolyzed linker species, and over-modified byproducts may co-elute unless purification is designed around the actual conjugate properties. Method selection frequently needs adjustment beyond a standard oligo HPLC workflow.
For many teams, the bottleneck is not synthesis alone but obtaining convincing LC-MS, purity, and integrity data that distinguish the intended conjugate from partially reacted or degraded material. We structure analytics around the decision points the project actually needs.
Our Peptide Oligonucleotide Conjugation Services
We provide modular support for peptide–oligonucleotide projects ranging from early feasibility studies to conjugates prepared for expanded biological evaluation. Service scope can include peptide selection, oligonucleotide handle design, site-specific conjugation, purification, analytical characterization, and route refinement for repeat synthesis.
 Conjugate Feasibility & Architecture Design
Conjugate Feasibility & Architecture Design
Capabilities include:
- Review of peptide class, oligonucleotide modality, and project objective before chemistry selection
- Recommendation of 5′, 3′, or internal attachment strategies based on function-sensitive regions
- Selection of orthogonal reactive handles for cleaner assembly
- Linker planning for stable, cleavable, or spacer-assisted formats
- Risk assessment covering activity retention, aggregation, and purification difficulty
Typical use cases:
Early-stage construct screening, rescue of underperforming uptake profiles, and design of defined peptide-linked formats for structure–activity evaluation
 Custom CPP and Targeting Peptide–Oligo Conjugation
Custom CPP and Targeting Peptide–Oligo Conjugation
Capabilities include:
- Conjugation of cell-penetrating, receptor-targeting, localization, or functional peptide sequences to ASO, siRNA, DNA, and RNA payloads
- Support for terminal and selected internal modification handles
- Alignment of peptide sequence design with charge, hydrophobicity, and downstream assay constraints
- Conjugate preparation for uptake, trafficking, and activity comparison studies
- Optional integration with related peptide RNA conjugation workflows when the payload is RNA-focused
Typical applications:
Intracellular delivery studies, receptor-directed uptake programs, localization-driven constructs, and custom research tools
 PMO, SSO, and Neutral-Backbone Conjugate Support
PMO, SSO, and Neutral-Backbone Conjugate Support
Capabilities include:
- Peptide conjugation strategies adapted to PMO and steric-blocking oligonucleotide formats
- Design support for splice-switching and intracellular trafficking studies
- Selection of chemistry compatible with backbone and deprotection limitations
- Route planning for constructs where direct synthesis is impractical and modular assembly is preferred
- Adjacent support for peptide-linked analog systems such as peptide-PNA conjugation when sequence recognition formats are being compared
Focus areas:
Steric-blocking constructs, CPP-assisted PMO programs, and research-stage conjugates requiring careful backbone-specific handling
 Site-Specific Linker & Handle Engineering
Site-Specific Linker & Handle Engineering
Capabilities include:
- Azide, alkyne, amine, thiol, maleimide, and other orthogonal handle planning
- Selection among amide, thioether, disulfide, and click-derived linkages
- Spacer optimization to reduce steric interference and improve chromatographic behavior
- Evaluation of stable versus cleavable linker concepts based on the intended use
- Troubleshooting of low conversion or unstable conjugation routes
Deliverables:
Recommended linker format, attachment map, chemistry rationale, and practical route notes for synthesis execution
 Purification, Deconvolution & Analytical Characterization
Purification, Deconvolution & Analytical Characterization
Capabilities include:
- RP-HPLC, ion-pair RP-HPLC, or orthogonal purification planning depending on conjugate properties
- LC-MS confirmation of expected mass and conjugation integrity
- Assessment of unconjugated payload, truncated species, and hydrolysis-related byproducts
- Purity reporting structured for project review and method comparison
- Comparative analysis across pilot batches or alternative conjugation routes
Typical outputs:
Chromatograms, deconvoluted mass data, peak assignment summaries, and route-specific analytical observations
 Route Refinement for Repeat Synthesis
Route Refinement for Repeat Synthesis
Capabilities include:
- Reproducibility-oriented optimization of reaction order, equivalents, and purification conditions
- Evaluation of whether on-column, post-synthetic, or staged assembly is the most robust route
- Batch planning for expanded in vitro and in vivo research studies
- Technical comparison against alternative conjugate classes such as PEG-conjugated oligonucleotides or cholesterol-conjugated oligonucleotides when peptide is not the only delivery option under consideration
Project value:
Greater process clarity, more consistent repeat batches, and cleaner decision-making when advancing or redirecting a conjugation strategy
Core Design Parameters for Peptide–Oligonucleotide Conjugates
In peptide oligonucleotide conjugation, construct performance depends on coordinated control of peptide function, oligonucleotide chemistry, conjugation site, linker behavior, and downstream analytical tractability. The table below summarizes the design variables that most directly influence feasibility, purity, and biological usefulness.
| Design Parameter | Typical Options | Why It Matters | Main Development Risk | Our Design Focus |
| Oligonucleotide Format | ASO, siRNA, PMO, SSO, DNA, RNA | Different oligo classes have different mechanism sensitivity, backbone compatibility, and conjugation tolerance | Loss of activity if the conjugation site or chemistry interferes with hybridization, RNAi loading, or steric-blocking function | Match conjugation strategy to the oligo mechanism and modification pattern |
| Peptide Type | CPP, targeting peptide, localization peptide, functional peptide | The peptide determines whether the conjugate is being used for uptake enhancement, targeting, intracellular routing, or assay utility | Poor uptake improvement, off-target interactions, aggregation, or excessive sequence complexity | Select peptide class according to the intended biological problem the conjugate needs to solve |
| Attachment Position | 5′ terminus, 3′ terminus, selected internal handle | Attachment position can strongly affect oligo accessibility and functional performance | Steric interference, reduced potency, or increased heterogeneity | Choose the least function-sensitive position while keeping the route synthetically practical |
| Linker Format | Stable spacer, cleavable linker, disulfide, triazole, amide, thioether | Linker choice affects stability, release concept, flexibility, and chromatographic behavior | Premature degradation, poor intracellular release logic, or difficult purification | Balance stability, mechanism relevance, and purification compatibility |
| Backbone / Sugar Chemistry | PO/PS patterns, 2′-OMe, 2′-F, MOE, LNA, PMO-compatible formats | Oligo chemistry determines how aggressive conjugation and deprotection conditions can be | Backbone damage, incomplete conversion, or incompatibility with the selected coupling route | Use chemistry-aware route design rather than a one-method-fits-all workflow |
| Purification Strategy | RP-HPLC, ion-pair RP-HPLC, orthogonal re-purification | Peptide and oligo domains change charge, hydrophobicity, and impurity profile simultaneously | Co-elution of unconjugated payload, truncated peptide species, or partially reacted product | Develop purification around the actual conjugate behavior instead of generic oligo release conditions |
Recommended Peptide–Oligonucleotide Conjugate Designs by Project Scenario
For most clients, the key question is not simply which conjugation chemistry exists, but which peptide–oligo design is more suitable for the actual project goal. The table below translates common development needs into practical construct planning logic.
| Project Scenario | Recommended Peptide Direction | Typical Oligo Types | Preferred Design Considerations | Key Risks to Evaluate |
| Need to Improve Cellular Uptake | Cell-penetrating peptides with controlled charge density | ASO, PMO, SSO, selected siRNA formats | Terminal attachment, spacer-assisted design, and sequence tuning to limit aggregation | High cationic content may complicate purification and increase non-specific interactions |
| Need More Selective Cell or Receptor Engagement | Targeting peptide matched to receptor or cell-surface binding objective | ASO, siRNA, DNA, RNA probes | Preserve the peptide binding motif while keeping the oligo accessible for activity | Reduced targeting performance if the conjugation position disrupts peptide recognition |
| Need Better Intracellular Routing or Endosomal Processing | Peptides with membrane-interacting or trafficking-support properties | ASO, PMO, SSO, selected RNA formats | Linker flexibility and peptide placement should be assessed together with the intended uptake model | Improved uptake without improved functional release can still lead to weak biological performance |
| Need Defined Research Tools for Tracking or Mechanistic Studies | Localization, affinity, or functional peptide motifs | DNA, RNA, ASO, labeled probe formats | Favor analytically clean, site-defined constructs with clear stoichiometry | Overengineering the construct may reduce interpretability in downstream assays |
| Need CPP-Linked PMO or Steric-Blocking Constructs | CPPs compatible with PMO or splice-switching workflows | PMO, SSO | Use backbone-compatible route design and avoid conditions that compromise neutral-backbone integrity | Route incompatibility, low conversion, and purification bottlenecks are common if the chemistry is not planned early |
| Need Multiple Variants for SAR or Feasibility Screening | Modular peptide panel with a common oligo scaffold | ASO, siRNA, PMO, DNA, RNA | Orthogonal handles and scalable post-synthetic assembly are usually preferred | Inconsistent construct quality can obscure true structure–activity conclusions |
Analytical Characterization, QC, and Typical Deliverables
In peptide oligonucleotide conjugation projects, analytical quality often determines whether the final material is genuinely decision-ready. Clients typically need more than nominal mass confirmation; they need evidence that the intended construct has been formed cleanly, that major impurities are understood, and that the batch is suitable for meaningful downstream evaluation.
| Analytical / QC Item | Typical Method | What It Confirms | Why Clients Care | Typical Deliverable |
| Identity Confirmation | LC-MS, high-resolution mass analysis, deconvolution review | The peptide and oligonucleotide are present in the intended conjugate mass | Confirms that the target construct was actually formed rather than an incomplete or misassigned product | Expected vs. observed mass summary with supporting spectra |
| Purity Assessment | RP-HPLC, ion-pair RP-HPLC, or orthogonal chromatographic method | Relative abundance of the main conjugate peak and separation of major impurities | Purity directly affects interpretability of uptake, trafficking, and activity studies | Chromatogram, purity estimate, and release summary |
| Impurity Profiling | LC-MS-supported peak assignment and route-specific impurity review | Presence of unconjugated oligo, truncated peptide species, linker hydrolysis products, or partially reacted intermediates | Helps distinguish a chemistry problem from a biology problem during project evaluation | Impurity interpretation notes and assigned peak summary |
| Conjugation Integrity | LC-MS before/after coupling comparison and targeted integrity review | Whether the final linkage remains intact through workup, purification, and storage handling | Critical when using disulfide, cleavable, or hydrolysis-sensitive linker concepts | Integrity assessment with route-specific observations |
| Repeat-Batch Comparability | Comparative HPLC and LC-MS review across repeat syntheses | Whether the route is reproducible enough for expanded studies | Essential when moving from a one-off feasibility build to repeat material supply | Batch comparison summary and analytical overlay review |
| Stability-Oriented Check | Defined-condition storage monitoring with periodic analytical follow-up | Short-term behavior of the conjugate under handling or storage conditions | Supports shipping, storage, and study planning decisions | Stability observations and handling recommendations |
Typical Workflow for a Peptide Oligonucleotide Conjugation Project
We start by reviewing the oligo type, peptide sequence, desired mechanism, and intended assay or model so that the conjugation plan is built around the project objective rather than around a default chemistry platform.
Orthogonal reactive groups, spacer length, and stable versus cleavable linkage concepts are selected with attention to activity retention, purification feasibility, and the expected biological readout.
The peptide and oligonucleotide components are synthesized or prepared in the most practical order for the chosen route, with protecting groups and reaction compatibility considered early to reduce downstream rework.
Coupling conditions are tuned for conversion, selectivity, and manageable impurity formation. Where needed, alternative reaction order or staged assembly is assessed to improve robustness.
Purification is matched to the conjugate properties, followed by LC-MS and chromatographic analysis to confirm identity, estimate purity, and flag any route-specific impurity patterns.
Final materials are supplied with the relevant analytical outputs and practical observations so your team can move more efficiently into uptake studies, activity assays, comparative screening, or repeat-batch planning.
Why Clients Choose Our Peptide–Oligonucleotide Conjugation Support
We evaluate peptide class, oligo mechanism, and the likely trafficking requirement together so that the final construct is designed to support the intended biology instead of merely producing a synthetically valid linkage.

Peptide-rich, hydrophobic, cationic, or backbone-sensitive projects often fail when handled as routine oligo modifications. Our workflow is built for constructs where sequence composition and purification behavior genuinely matter.
We support decisions around terminal versus internal attachment, orthogonal handle pairing, and staged assembly so that low conversion, scrambling, or poor peak resolution can be addressed at the design level.
Instead of stopping at nominal mass confirmation, we structure purity review, impurity interpretation, and repeat-batch comparison around the questions project teams usually need answered before the next study begins.
Research and Development Applications of Peptide–Oligonucleotide Conjugates
Cell-Penetrating Delivery Studies
- Covalent attachment of CPPs to ASO, PMO, or siRNA constructs for uptake and trafficking evaluation.
- Comparison of peptide sequence variants to improve intracellular exposure in difficult cell models.
- Screening of linker and attachment position effects on functional delivery.
Receptor-Targeted Oligonucleotide Programs
- Use of targeting peptides to explore cell-selective or tissue-biased uptake concepts.
- Defined conjugate formats for head-to-head comparison against non-peptide delivery strategies.
- Support for receptor-binding peptide selection where covalent format control is important.
Splice-Switching and Steric-Blocking Constructs
- Peptide-assisted PMO and SSO formats for intracellular trafficking and exon-level assay workflows.
- Construct design that prioritizes activity retention while improving cellular access.
- Useful for programs where unconjugated steric-blocking oligos underperform in uptake-sensitive systems.
Intracellular Imaging and Tracking Tools
- Peptide-linked oligo constructs prepared for localization, uptake, or pathway-tracking studies.
- Integration with additional labels or handles when the analytical plan supports them.
- Suitable for mechanistic work on internalization and compartmental routing.
Surface Capture, Assembly, and Biosystems Engineering
- Peptide–DNA or peptide–RNA conjugates for controlled assembly, immobilization, or binding studies.
- Defined constructs for hybrid materials, biointerfaces, and sequence-programmable systems.
- Useful when peptide functionality and oligo addressability need to coexist in one molecule.
Comparative Conjugate Platform Evaluation
- Side-by-side testing of peptide-linked constructs against lipid, carbohydrate, or polymer-linked alternatives.
- Support for platform down-selection based on uptake, tractability, and analytical clarity.
- Practical for teams deciding which conjugate class deserves deeper development effort.
Discuss Your Peptide Oligonucleotide Conjugation Strategy with Our Team
Whether you are evaluating a first peptide-linked oligo concept, troubleshooting a low-yield conjugation route, or comparing peptide delivery against other oligonucleotide conjugate formats, we can help build a project plan around your sequence, chemistry, and analytical priorities.
Our support is structured to reduce avoidable iteration at the interface between peptide design, oligonucleotide modification, conjugation chemistry, and purification. The result is a clearer path from concept to interpretable material for research and preclinical studies. Contact our scientific team to discuss your peptide oligonucleotide conjugation project, or share your sequence and target construct format for a tailored feasibility review.
Frequently Asked Questions (FAQ)
Different peptides can change how the conjugate behaves in solution, including folding and interactions with other molecules. Choosing the right peptide helps improve experimental reliability.
By using site-specific chemistry and careful reaction conditions, researchers can limit the number of peptides on each oligonucleotide, ensuring more uniform conjugates.
The position of peptides affects how the conjugate folds and interacts with other molecules. Proper placement avoids interference and keeps experiments consistent.

