Antibody Cell Conjugation (ACC)
Non-Genetic Cell Surface EngineeringTargeted Immune Cell RedirectionCustom Antibody-Cell Conjugation for Research Programs
Accelerate cell engineering studies with custom antibody-cell conjugation services designed for biotech, pharmaceutical, and academic research teams that need antibodies displayed on the surface of live cells without relying on stable genetic modification. Antibody-cell conjugation, often referred to as ACC, enables controlled installation of targeting antibodies onto immune or other living cells to support rapid target-switching, cell-surface functionalization, and mechanism-focused evaluation in complex research workflows.
We support antibody-cell conjugation projects involving NK cells, γδ T cells, CIK cells, primary T cells, and other compatible cell types by integrating antibody selection, surface chemistry strategy, conjugation development, and post-conjugation characterization into one coordinated workflow. Depending on your project goals, we can evaluate chemoenzymatic glycoengineering, DNA-directed assembly, bioorthogonal click approaches, or other live-cell-compatible chemistries to balance antibody loading, cell viability, wash stability, and functional performance.
What Problems Can Antibody-Cell Conjugation Solve?
Antibody-cell conjugation is especially valuable when a research team needs to give cells new targeting behavior but wants to avoid the longer timelines, process complexity, or construct constraints associated with permanent genetic engineering. In practice, this approach helps solve several common project bottlenecks: rapidly comparing multiple antibodies on the same cell chassis, improving target-cell engagement in co-culture systems, exploring non-genetic cell-surface engineering strategies, and testing whether a specific antibody can redirect cell interactions without compromising baseline phenotype. For many discovery programs, ACC offers a modular route to functionalize live cells while keeping close control over conjugation density, antibody orientation, residual free antibody, and downstream assay compatibility.
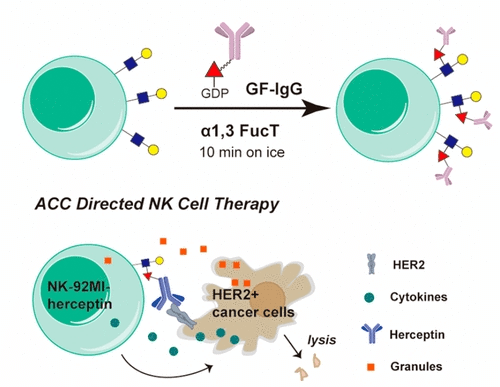 Fig 1. Enzymatic antibody-cell conjugates (Li, 2018)
Fig 1. Enzymatic antibody-cell conjugates (Li, 2018)
Key Technical Challenges in Antibody-Cell Conjugation Projects
Live-cell modification must be performed under conditions that limit membrane damage, aggregation, and unintended activation. We select reaction conditions, buffers, incubation windows, and cleanup steps with the goal of maintaining viability and preserving the phenotype needed for downstream binding, trafficking, or cytotoxicity assays.
Too little antibody can weaken target engagement, while excessive or poorly oriented loading can reduce accessibility or alter cell behavior. We evaluate conjugation chemistry, linker architecture, and loading level to improve functional presentation of the antibody on the cell surface.
The right strategy depends on antibody format, available reactive groups, cell-surface composition, and the required persistence of surface display. We compare feasible routes—including enzymatic transfer, click chemistry, DNA-mediated assembly, and other surface-reactive approaches—before committing to a development path.
Residual unconjugated antibody can complicate interpretation of binding or co-culture data. Our workflows include cleanup, analytical confirmation, and batch-to-batch comparison so that conjugated materials are suitable for consistent research use rather than one-off proof-of-concept preparation.
 Controlled antibody-cell conjugation can help research teams improve target engagement while maintaining live-cell compatibility and analytical consistency.
Controlled antibody-cell conjugation can help research teams improve target engagement while maintaining live-cell compatibility and analytical consistency.
Our Antibody-Cell Conjugation Services
We provide modular ACC development support for teams that need a practical, technically grounded path from feasibility to research-ready conjugated cells. Services can be configured around customer-supplied antibodies and cells or around a broader custom bioconjugation service package that covers design, chemistry development, analytical verification, and functional follow-up.
 Feasibility Assessment & Conjugation Design
Feasibility Assessment & Conjugation Design
Capabilities include:
- Review of target biology, intended cell type, and assay objective before chemistry selection
- Evaluation of full-length antibodies, fragments, or pre-functionalized antibody materials
- Selection of suitable cell-surface handles and antibody modification routes
- Antibody consumption planning for early feasibility versus expanded study needs
- Risk assessment for steric hindrance, off-target adsorption, and phenotype disturbance
- Integration with broader antibody conjugation services when upstream antibody derivatization is required
Typical use cases:
Early-stage cell engineering design, target-switching studies, and evaluation of whether non-genetic surface functionalization is the right fit for a specific research program
 Live-Cell Conjugation Method Development
Live-Cell Conjugation Method Development
Capabilities include:
- Screening of chemoenzymatic glycoengineering, DNA-directed assembly, and live-cell-compatible click routes
- Strategy selection informed by bioorthogonal reactions and other mild conjugation approaches
- Assessment of enzyme-enabled transfer or related enzymatic crosslinking services when cell compatibility is critical
- Optimization of reaction time, stoichiometry, linker length, and cleanup steps
- Development of workflows for NK cells, γδ T cells, CIK cells, primary T cells, and other compatible cell types
- Adaptation to customer antibodies, antibody fragments, or pre-modified intermediates
Focus areas:
Fast feasibility screening, improved surface display stability, and method selection based on live-cell compatibility rather than one-chemistry-fits-all assumptions
 Analytical Characterization & Functional Validation
Analytical Characterization & Functional Validation
Capabilities include:
- Flow cytometry-based confirmation of antibody loading and target-binding retention
- Evaluation of post-conjugation viability, phenotype markers, and aggregation tendency
- Residual free-antibody assessment after purification or wash steps
- Microscopy or orthogonal binding assays to confirm surface presentation
- Functional readouts tailored to project goals, such as target-cell binding or co-culture activity
- Comparative datasets to identify the best-performing chemistry or loading window
Deliverables:
Data packages that clarify whether the final conjugated cells meet the handling, assay, and reproducibility requirements of your study
 Process Optimization for Repeated Research Batches
Process Optimization for Repeated Research Batches
Capabilities include:
- Translation of initial feasibility work into repeatable batch workflows
- Definition of critical process parameters affecting antibody loading and cell recovery
- Side-by-side batch comparison to reduce variability across repeated studies
- Guidance on material handling, storage windows, and assay scheduling after conjugation
- Technical documentation to support internal transfer or future expansion of the workflow
- Coordination with adjacent protein conjugation services when accessory proteins or ligands are involved
Typical outputs:
Standardized preparation records, recommended operating ranges, and practical decision guidance for repeated ACC studies
Key Design Parameters for Antibody-Cell Conjugation
Successful ACC development depends on choosing a chemistry and process that fit both the antibody and the living cell. The table below summarizes the design variables that typically have the greatest impact on usability, stability, and downstream assay performance.
| Design Parameter | Common Options | What to Evaluate | Impact on Project Outcome | Typical Decision Point |
| Cell Chassis | NK cells, γδ T cells, CIK cells, primary T cells, other compatible live cells | Baseline phenotype, membrane composition, activation sensitivity, and available cell numbers | Influences chemistry tolerance, recovery, and functional assay design | Selected at project start based on biological objective and handling constraints |
| Antibody Format | Full-length IgG, fragments, or pre-functionalized antibody intermediates | Reactive group availability, size, orientation needs, and target-binding retention | Affects conjugation route, steric accessibility, and loading efficiency | Defined during feasibility and upstream material review |
| Conjugation Strategy | Chemoenzymatic transfer, click-mediated attachment, DNA-directed assembly, or other surface-reactive methods | Cell compatibility, reaction speed, positional control, and wash stability | Determines the balance between mild processing and robust surface display | Compared in small-scale screens before process lock-in |
| Antibody Loading Level | Low, medium, or high surface density windows established experimentally | Binding improvement versus phenotype disturbance, aggregation, and cell recovery | Directly affects target engagement and assay reproducibility | Optimized after first-pass conjugation results are reviewed |
| Linker or Spacer Design | Short, intermediate, or more flexible architectures depending on chemistry | Antigen accessibility, membrane proximity effects, and conjugate stability | Can improve functional presentation while reducing steric masking | Refined when initial loading is acceptable but functional performance lags |
| Post-Conjugation Handling | Immediate use, short hold, wash-intensive workflows, or repeated assay preparation | Signal retention, free-antibody removal, and stability during handling | Shapes the final operating procedure and scheduling flexibility | Confirmed during validation of the intended downstream assay workflow |
Antibody-Cell Conjugation Strategies and Method Selection
Different ACC methods solve different project problems. Rather than forcing every program into a single chemistry, we compare options according to cell compatibility, process complexity, persistence of surface display, and the type of analytical evidence required afterward.
| Conjugation Strategy | Technical Basis | When It Is Often Considered | Key Practical Considerations |
| Chemoenzymatic Glycoengineering | Enzyme-enabled transfer of antibody-bearing substrates onto suitable cell-surface glycans | Projects prioritizing mild conditions and efficient modification of living cells | Requires compatible enzyme and substrate design, but can offer strong live-cell compatibility and fast processing |
| Metabolic Glycoengineering + Click Chemistry | Installation of reactive handles on the cell surface followed by bioorthogonal coupling to modified antibodies | Programs needing modular chemistry selection and flexible handle pairing | Handle density, incubation time, and click partner choice influence loading efficiency and phenotype preservation |
| DNA-Directed Assembly | Complementary oligonucleotide strands are introduced onto antibody and cell surfaces, then hybridized | Comparative screening programs and applications requiring modular exchange of targeting antibodies | Sequence design, hybridization stability, and additional modification steps must be managed carefully |
| Surface-Reactive Covalent Coupling | Direct coupling to accessible functional groups on the cell surface under controlled live-cell conditions | Feasibility studies where rapid implementation is important and materials are limited | Useful for screening, but requires careful control of specificity, surface accessibility, and cell stress |
| Hybrid or Stepwise Workflows | Combined approaches that separate antibody modification, cell preparation, conjugation, and cleanup into defined modules | More complex programs that need higher control over intermediate quality | Often preferred when the project must compare multiple antibodies, linker designs, or cell types side by side |
| Assay-Aligned Method Optimization | Final process tuned around the downstream readout rather than around conjugation efficiency alone | Binding, co-culture, imaging, or functional studies where assay compatibility is critical | Prevents selection of a chemistry that looks strong analytically but performs poorly in the intended biological workflow |
Analytical Characterization and QC Framework for ACC Materials
Research-grade antibody-cell conjugates still require disciplined characterization. Our analytical framework is designed to answer the questions that matter most to project teams: Did the antibody load successfully, does it remain functional, are unconjugated materials sufficiently controlled, and do repeated batches behave similarly enough for comparative studies?
| Analytical Category | Typical Method | Purpose in the Project | Data Delivered |
| Cell Viability & Recovery | Viability dye analysis, cell counting, and recovery comparison before versus after conjugation | Confirms that the selected chemistry does not excessively compromise the live-cell preparation | Viability percentages, recovery tables, and batch comparison summaries |
| Antibody Loading Confirmation | Flow cytometry, fluorescence-based detection, or orthogonal surface-binding assays | Verifies that antibodies are displayed on the cell surface at useful levels | Loading histograms, positive population percentages, and relative signal comparisons |
| Target-Binding Retention | Antigen-binding assays, competitive binding studies, or microscopy-based assessment | Demonstrates that antibody recognition is retained after conjugation | Binding curves, comparative target-cell interaction data, or imaging outputs |
| Residual Free Antibody Assessment | Wash monitoring, supernatant analysis, and signal comparison before and after cleanup | Reduces the risk of misattributing free antibody effects to conjugated cells | Cleanup efficiency records and supporting analytical notes |
| Phenotype Maintenance | Marker panel assessment relevant to the chosen cell type | Confirms that critical surface markers or activation state are not unintentionally altered | Phenotype comparison tables and representative plots |
| Functional Readout | Target-cell binding, co-culture response, signaling, or other project-specific activity assays | Determines whether the conjugated cells perform better than relevant controls in the intended assay | Functional comparison dataset and interpretation summary |
| Batch Consistency | Side-by-side comparison of repeated preparations using the same workflow | Supports reproducible use of the method across multiple studies | Lot comparison reports, analytical overlays, and recommended operating ranges |
| Technical Documentation | Structured reporting of materials, process parameters, and analytical outputs | Facilitates internal review, knowledge transfer, and follow-on optimization | Project report, raw-data summary, and decision-focused recommendations |
A Practical Workflow for Antibody-Cell Conjugation Programs
We begin by reviewing cell type, target antigen, antibody format, required assay readouts, and available starting materials. This step clarifies the real objective of the program—rapid screening, deeper method development, or repeated-batch preparation—and prevents unnecessary chemistry work.
Candidate routes are compared at small scale to determine which chemistry is compatible with the live cells and antibody while still delivering useful loading. Early screening focuses on viability, loading efficiency, surface accessibility, and cleanup practicality.
Once a route is chosen, we prepare the antibody intermediate and cell surface in a controlled sequence. This stage is where linker choice, reagent stoichiometry, cell handling conditions, and reaction timing are refined to reduce stress on the cell material.
Antibodies are coupled to the live-cell surface under defined conditions, followed by washing or purification steps to reduce free antibody background. Initial retention studies help determine whether the conjugates remain suitable for the intended handling and assay workflow.
We verify antibody loading, target-binding retention, cell recovery, and phenotype integrity, then connect those results to the relevant biological readout. This prevents decision-making based only on signal intensity while ignoring whether the conjugated cells are actually usable.
Final output includes a structured review of what worked, what constrained the process, and which parameters should be fixed or optimized next. Where required, we translate the best-performing conditions into a repeatable workflow for subsequent studies.
Why Teams Choose Our Antibody-Cell Conjugation Platform
We do not assume that one conjugation route fits every cell type and antibody. Projects are designed around the practical trade-offs between loading efficiency, cell compatibility, cleanup burden, and downstream assay needs.

ACC succeeds only when the cells remain usable. Our development logic prioritizes reaction conditions and processing windows that protect cell recovery, viability, and phenotype instead of maximizing loading at any cost.
Conjugation efficiency alone is not enough. We combine loading analysis with binding, cleanup, phenotype, and functional readouts so project teams can judge whether a method is analytically attractive and biologically useful.
From a one-antibody feasibility check to repeated study batches with detailed reporting, we can adapt the scope of work to fit exploratory screening, mechanism studies, or more structured method development programs.
Research Applications of Antibody-Cell Conjugation
Immune Cell Retargeting Studies
- Exploration of whether surface-installed antibodies can redirect NK, CIK, or T-cell interactions toward defined target cells.
- Comparative studies using different antibodies on the same cell chassis.
- Assessment of target-binding gain without stable gene transfer.
Cell-Surface Engineering Research
- Installation of new recognition elements on living cells for functional surface remodeling.
- Evaluation of how surface chemistry changes cell-cell communication.
- Mechanistic studies on receptor access and membrane presentation.
Co-Culture and Target Engagement Assays
- Improvement of target-cell engagement in mixed-cell systems.
- Comparison of antibody density, linker design, or conjugation route in the final assay context.
- Reduction of uncertainty when selecting a chemistry for more advanced follow-up work.
Antibody Screening on a Common Cell Platform
- Rapid switching between target-binding antibodies without rebuilding a genetic construct each time.
- Side-by-side evaluation of antibody clones, fragments, or modified variants.
- More efficient ranking of candidates before deeper development investment.
Mechanism-of-Action Studies
- Investigation of how antibody-armed cells interact with target cells at the binding and signaling level.
- Linking surface loading and target recognition to downstream functional response.
- Use of analytical controls to separate conjugate-driven effects from free-antibody artifacts.
Custom Proof-of-Concept Programs
- Tailored studies that combine antibody preparation, cell conjugation, analytics, and functional testing.
- Feasibility evaluation for uncommon antibodies or specialized cell systems.
- Structured next-step recommendations for future optimization rounds.
Move Your Antibody-Cell Conjugation Project Forward with a Practical Development Partner
Whether you are exploring a non-genetic cell-surface engineering concept, comparing multiple antibodies on a single live-cell platform, or building a more repeatable ACC workflow for ongoing studies, our team can help translate the idea into a technically workable process.
We combine conjugation design, live-cell process development, analytical confirmation, and functional readout alignment so your team can make decisions based on usable evidence rather than chemistry alone.Contact our scientific team to discuss your antibody-cell conjugation goals and define a project scope tailored to your cells, antibodies, and assay requirements.
Frequently Asked Questions (FAQ)
Unlike solution-phase conjugation or solid-phase immobilization, Antibody Cell Conjugation directly couples antibodies onto intact cell surfaces. This enables real-time studies of cell-molecule interactions, receptor mapping, and surface functionalization under physiological conditions.
Different conjugation strategies—such as amine coupling, thiol-maleimide reactions, or click chemistry—can alter how antibodies orient on the cell surface. Selecting a suitable method ensures proper antigen recognition while minimizing steric interference or membrane disruption.
Factors such as cell type, membrane composition, and surface protein abundance significantly affect conjugation outcomes. Pre-conditioning steps, including washing, activation, or fixation, are optimized to maintain consistent binding efficiency and cell compatibility.
Consistency is verified through quantitative fluorescence analysis, binding ratio determination, and inter-batch comparison. Our workflow integrates standardized conjugation protocols and analytical checkpoints to achieve reproducible and comparable data outputs.

