Lipid Conjugation
Membrane Anchoring & Delivery OptimizationCustom Lipid ConjugatesLipid Conjugation for Oligonucleotides, Peptides, Proteins & Nanocarriers
Accelerate lipid-enabled molecular design with custom lipid conjugation services built for research teams working on oligonucleotides, peptides, proteins, small molecules, liposomes, and lipid nanoparticle-related systems. Lipid conjugation introduces carefully selected hydrophobic motifs—such as cholesterol, fatty acids, phospholipids, or reactive PEG-lipids—to improve membrane interaction, serum association, carrier compatibility, and delivery behavior while keeping the underlying payload strategy aligned with your project goals.
We support both direct biomolecule lipidation and carrier-oriented surface functionalization, covering lipid–oligonucleotide, lipid–peptide, lipid–protein, and lipid-enabled liposome workflows. Projects can be designed around attachment site, linker behavior, conjugation stoichiometry, and downstream analytical requirements, whether you need a simple hydrophobic anchor, a membrane-inserting construct, or a reproducible route for ligand display on preformed lipid systems. Related programs can also be integrated with custom bioconjugation services when broader conjugation design is required.
How Lipid Conjugation Solves Delivery and Formulation Bottlenecks
Many promising biomolecules underperform not because of weak target binding, but because they lack the physical properties needed for practical delivery and formulation. Unmodified oligonucleotides may show limited cellular or carrier association, peptides can clear quickly or lose activity in solution, proteins may require a membrane anchor for surface presentation, and liposomal systems often need controlled ligand display rather than non-specific adsorption. Lipid conjugation addresses these problems by introducing a defined lipophilic element that can increase membrane affinity, improve association with serum components, support self-assembly or carrier loading, and enable more stable integration into lipid bilayers. The key is not simply adding a lipid, but matching lipid class, linker length, and attachment site to the payload so that uptake, stability, and function improve together rather than at each other's expense.
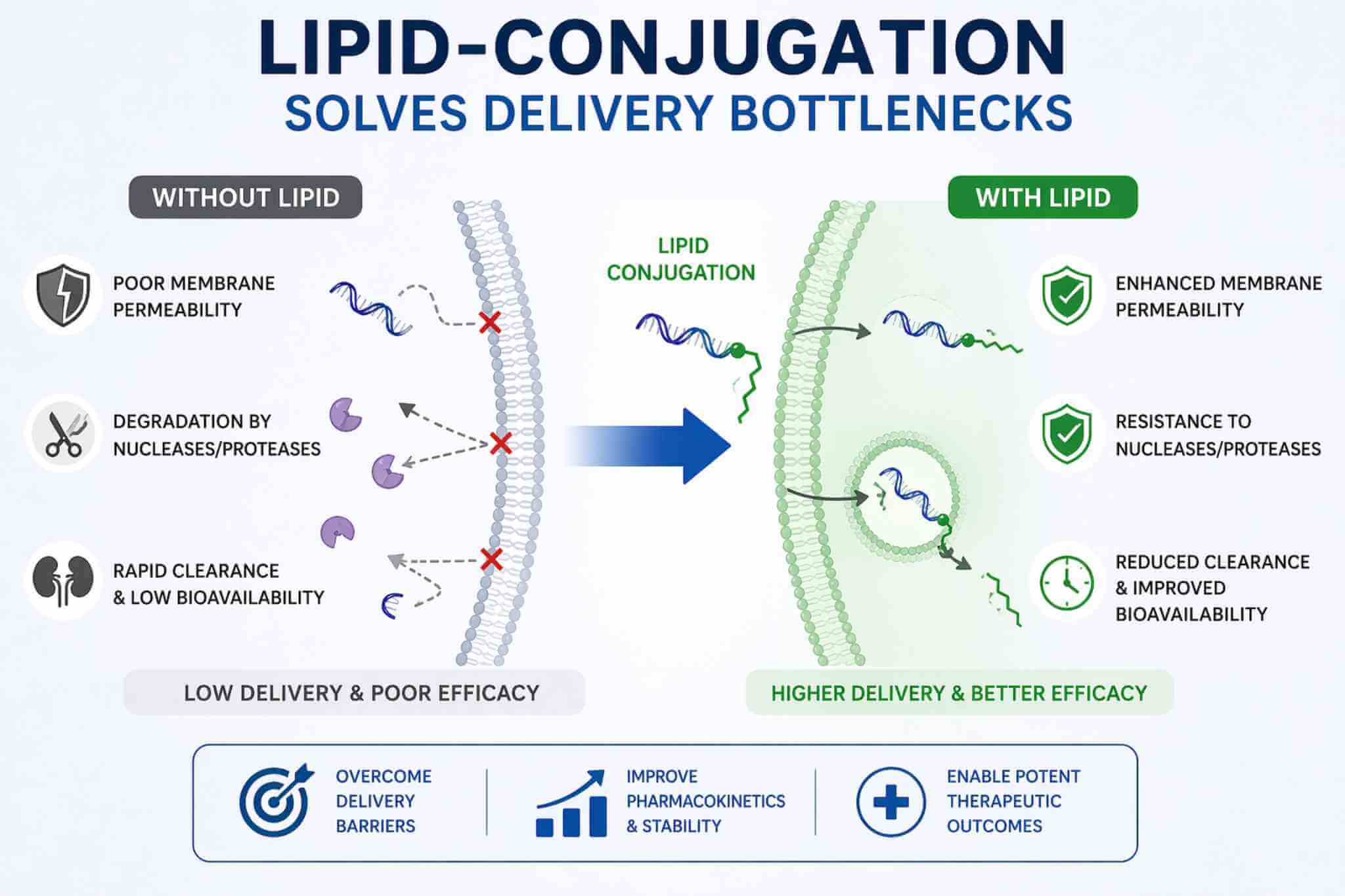 Concept image showing how lipid conjugation improves membrane anchoring, carrier integration, and delivery-oriented molecular design for oligonucleotides, peptides, proteins, and liposomal systems.
Concept image showing how lipid conjugation improves membrane anchoring, carrier integration, and delivery-oriented molecular design for oligonucleotides, peptides, proteins, and liposomal systems.Common Development Challenges Lipid Conjugation Can Address
Molecules that are potent in biochemical assays often underperform once delivery, membrane contact, or tissue exposure becomes limiting. We help select cholesterol, fatty acid, phospholipid, or PEG-lipid formats that are better matched to the intended uptake mechanism, membrane environment, and carrier architecture.
A lipid can improve hydrophobic interaction while simultaneously disrupting hybridization, receptor binding, enzymatic recognition, or folding if the attachment site is poorly chosen. Our design workflows emphasize site control and linker spacing so lipidation supports delivery performance without masking the functional region of the payload.
Lipidated constructs can show new behaviors during synthesis, handling, and storage, including self-association, broad chromatographic peaks, or difficult removal of free lipid and partially modified species. We build purification and analytical plans around the expected amphiphilic behavior of each conjugate instead of treating it like a standard unconjugated biomolecule.
For liposome and related lipid carrier projects, inconsistent surface loading can directly affect targeting, colloidal behavior, and batch reproducibility. We support controlled attachment strategies, post-insertion or prefunctionalized lipid routes, and orthogonal analysis to verify conjugation efficiency, free ligand content, and carrier-level quality attributes.
Our Lipid Conjugation Services
We provide development-focused lipid conjugation support across biomolecule classes and carrier systems, combining chemistry selection, attachment-site planning, purification strategy, and analytical confirmation. Service scope can be adapted for early feasibility studies, lead optimization, comparative linker screening, or reproducible batch preparation for downstream research workflows.
 Lipid–Oligonucleotide Conjugation
Lipid–Oligonucleotide Conjugation
Capabilities include:
- Conjugation of cholesterol, fatty acid, phospholipid, tocopherol-related, or custom lipid motifs to siRNA, ASO, aptamer, DNA, and RNA constructs
- Support for terminal or internal attachment strategies depending on sequence format and functional constraints
- Linker selection to balance hydrophobicity, serum association, and payload accessibility
- Compatibility with common oligonucleotide modification patterns and screening-stage structure optimization
- HPLC and LC-MS confirmation of identity, purity, and conjugation integrity
- Natural extension into cholesterol-conjugated oligonucleotide and broader oligonucleotide bioconjugation programs
Typical applications:
Delivery-oriented oligonucleotide optimization, membrane-anchored probes, uptake studies, and carrier-compatible nucleic acid design
 Lipid–Peptide and Lipid–Protein Conjugation
Lipid–Peptide and Lipid–Protein Conjugation
Capabilities include:
- Fatty acid, cholesterol, and phospholipid conjugation for peptides, proteins, binders, enzymes, and related biologically active constructs
- Site-directed strategies based on free amines, thiols, engineered handles, or orthogonal click-compatible groups
- Design support for membrane anchoring, half-life-oriented albumin association, or amphiphilic self-assembly studies
- Attention to lipidation position to reduce disruption of folding, binding motifs, or catalytic regions
- Purification planning for hydrophobicity-shifted products and free-lipid removal
- Optional alignment with cholesterol-conjugated peptide and protein conjugation services where appropriate
Typical applications:
Membrane-active peptides, lipidated binders, protein display studies, and conjugates requiring improved bilayer interaction or controlled amphiphilicity
 Liposome and PEG-Lipid Surface Functionalization
Liposome and PEG-Lipid Surface Functionalization
Capabilities include:
- Conjugation of peptides, proteins, antibodies, oligonucleotides, and small ligands to reactive lipids or preformed lipid carriers
- Workflows based on DSPE-PEG-maleimide, NHS-reactive lipids, azide/alkyne pairs, and related surface-coupling handles
- Evaluation of direct coupling versus post-insertion strategies for ligand presentation on bilayer systems
- Support for liposome decoration, membrane anchoring studies, and targeted carrier engineering
- Orthogonal analysis of free ligand, conjugation efficiency, particle size, and colloidal behavior
- Highly relevant for peptide liposome conjugation, protein liposome conjugation, and oligonucleotide-loaded lipid nanoparticle projects
Focus areas:
Controlled ligand display, reproducible surface chemistry, and carrier-ready conjugation routes for lipid-based systems
 Lipid Linker Design, Purification, and Analytical Support
Lipid Linker Design, Purification, and Analytical Support
Capabilities include:
- Comparative evaluation of cleavable versus stable linkers and spacer length effects
- Purification route design for amphiphilic conjugates, including removal of unconjugated biomolecule and excess lipid reagent
- Identity confirmation by LC-MS and purity analysis by HPLC or UPLC
- Conjugation ratio assessment and free-species monitoring
- Particle characterization for liposome-related systems where required
- Technical reporting packages suited to internal R&D decision-making and method transfer discussions
Deliverables:
Chromatograms, mass confirmation, conjugation summary data, and project-specific technical recommendations for next-step optimization
Payload × Lipid Matching Table
Different payload classes respond very differently to lipid modification. In practice, the right lipid choice depends on whether the goal is membrane anchoring, improved carrier compatibility, controlled surface presentation, or a better balance between hydrophobicity and functional performance. This table helps map common payload types to the lipid formats most often considered during early project design.
| Payload Type | Common Lipid Options | Typical Attachment Site / Route | Primary Project Goal | Key Development Concern |
| Oligonucleotides | Cholesterol, fatty acids, phospholipids, tocopherol-related lipids | 5′ or 3′ terminal modification; pre-synthetic incorporation; post-synthetic coupling | Improve membrane interaction, uptake behavior, or carrier compatibility | Preserving hybridization, sequence function, and manageable purification |
| Peptides | Fatty acids, cholesterol, phospholipid anchors | N-terminus, C-terminus, lysine side chain, cysteine-directed attachment | Support membrane anchoring, amphiphilic behavior, or self-assembly studies | Avoiding activity loss, aggregation, or excessive hydrophobicity |
| Proteins | Fatty acids, cholesterol, phospholipids, reactive PEG-lipids | Lysine coupling, cysteine-selective chemistry, engineered handle modification | Enable surface display, membrane interaction, or controlled carrier association | Maintaining folding, binding activity, and defined modification level |
| Small Molecules / Probes | Cholesterol, fatty acids, custom hydrophobic anchors | Amide coupling, click-based linkage, custom handle integration | Create membrane-localized probes or lipid-compatible functional molecules | Maintaining recognition properties and structural clarity after conjugation |
| Liposome Surface Ligands | PEG-lipids, phospholipid anchors, preconjugated ligand-lipid intermediates | Reactive PEG-lipid coupling or post-insertion into preformed liposomes | Achieve controlled ligand presentation on bilayer surfaces | Ligand density, free ligand removal, and batch reproducibility |
| Lipid Nanocarrier-Associated Biomolecules | PEG-lipids, phospholipids, custom surface anchors | Surface-coupled integration, anchor-mediated insertion, or reactive lipid intermediates | Improve carrier functionality without compromising colloidal behavior | Balancing presentation efficiency with particle stability and analytical visibility |
Conjugation Chemistry / Strategy Table
Lipid conjugation can be achieved through several chemistry routes, but the best option depends on the payload, available functional groups, required site control, and whether the final product is a discrete molecular conjugate or a lipid-surface-functionalized system. This table summarizes the most commonly used conjugation strategies and the situations in which they are most useful.
| Chemistry / Strategy | Typical Reactive Pair | Best Suited For | Main Strength | Main Watchpoint |
| Amide Coupling | Amine + NHS ester / activated carboxyl group | Peptides, proteins, small molecules, selected lipid intermediates | Stable covalent linkage and broad synthetic accessibility | May produce heterogeneous modification if multiple amines are available |
| Thiol–Maleimide Conjugation | Thiol + maleimide-functional lipid or PEG-lipid | Cysteine-containing peptides, proteins, and liposome surface ligands | Useful for relatively controlled site-selective attachment | Requires careful handling of thiol availability and reaction conditions |
| Click Chemistry | Azide + alkyne or strain-promoted click pair | Sensitive biomolecules, modular assembly, orthogonal conjugation workflows | High selectivity and compatibility with multifunctional systems | Requires prior installation of compatible handles on both components |
| Pre-Synthetic Lipid Incorporation | Lipid-bearing building block introduced during synthesis | Oligonucleotides and peptides requiring precise positional control | Clear site definition and streamlined stoichiometry control | Less flexible once the synthesis design has been fixed |
| Post-Synthetic Covalent Coupling | Preformed payload reacted with activated lipid reagent | Rapid feasibility studies and comparative lipid screening | Flexible route for testing multiple lipid types without rebuilding the payload | Purification can become more challenging for amphiphilic products |
| Post-Insertion Strategy | Preconjugated lipid-ligand inserted into preformed lipid system | Liposomes and related lipid carrier surface-functionalization projects | Useful for formulation-stage ligand display optimization | Insertion efficiency and surface density should be verified analytically |
Linker Strategy Table
In lipid conjugation, the linker is often just as important as the lipid itself. Linker length, flexibility, hydrophilicity, and stability can determine whether the lipid improves delivery behavior without masking the active region of the payload or creating new formulation problems. The table below compares the linker strategies most often evaluated during development.
| Linker Type | Typical Behavior | Best Suited For | Main Advantage | Development Trade-Off |
| Direct Linkage | Minimal spacing between payload and lipid | Compact conjugates where steric separation is not critical | Simple architecture and reduced synthetic complexity | Can increase steric interference or reduce payload accessibility |
| Short Spacer Linker | Provides modest distance while keeping the construct compact | Peptides, probes, and selected oligonucleotide conjugates | Useful compromise between simplicity and functional separation | May still be insufficient when the active region is highly structure-sensitive |
| PEG Spacer | Hydrophilic and flexible spacing between lipid and payload | Liposome surface ligands, proteins, and sterically sensitive payloads | Improves exposure of the payload away from the lipid surface or bilayer | PEG length can affect surface density, mobility, and overall construct behavior |
| Non-Cleavable Linker | Maintains stable attachment throughout handling and testing | Projects prioritizing persistent membrane anchoring or surface display | High structural stability and clearer analytical interpretation | Not ideal when payload release or environment-responsive behavior is desired |
| Cleavable Linker | Designed to break under defined chemical or biological conditions | Projects requiring conditional release after delivery or localization | Adds an extra layer of functional control to the conjugate design | Requires more validation of cleavage behavior and storage stability |
| Hydrophilic / Solubilizing Linker | Reduces excessive hydrophobic clustering around the lipidated construct | Highly hydrophobic conjugates prone to aggregation or poor recovery | Can improve handling, purification, and solution behavior | May reduce the apparent membrane-driving effect of the lipid if overextended |
Analytical & QC Table
For lipid conjugates, a successful reaction is only part of the answer. Research teams usually need analytical evidence that the final material is structurally correct, sufficiently pure, appropriately modified, and reproducible enough for downstream studies. This table highlights the core analytical and QC categories that are most relevant to lipid-conjugated molecules and lipid-based systems.
| Analytical Item | Common Method | What It Confirms | Why It Matters |
| Identity Confirmation | LC-MS, HRMS, or MALDI-TOF | Correct molecular structure and expected lipid attachment | Verifies that the intended conjugate was formed rather than a related impurity or incomplete product |
| Purity Assessment | HPLC, UPLC, SEC, or orthogonal chromatographic methods | Relative proportion of target conjugate versus byproducts | Helps determine whether the material is clean enough for meaningful downstream work |
| Free Lipid / Free Payload Monitoring | Targeted chromatographic separation or orthogonal impurity analysis | Residual unconjugated lipid reagent and unconjugated biomolecule | Free species can distort uptake, formulation, and bioassay readouts |
| Conjugation Ratio / Modification Level | LC-based quantitation, mass distribution analysis, UV-based comparison, loading assay | Degree of lipidation or ligand density on the final construct | Critical for reproducibility and for comparing structure–activity results across variants |
| Particle Characterization | Dynamic light scattering, zeta potential, and formulation-level particle analysis | Size, PDI, surface charge, and colloidal consistency of lipid systems | Essential when the project involves liposomes, PEG-lipid assemblies, or carrier-associated biomolecules |
| Stability Monitoring | Time-course HPLC/LC-MS review under defined storage or handling conditions | Degradation, deconjugation, aggregation, or formulation drift over time | Shows whether the material can be stored and used consistently in planned experiments |
| Batch Comparability | Cross-batch chromatographic, mass-based, and particle-level comparison | Consistency between repeat preparations | Important when moving from exploratory screening to reproducible project execution |
Our Lipid Conjugation Workflow
We begin by reviewing payload class, available functional handles, intended delivery context, solubility limits, and any prior structure–activity observations. This helps define whether the project is best served by direct lipidation, reactive PEG-lipid coupling, or surface functionalization of a lipid carrier.
We shortlist lipid class, linker type, and candidate attachment sites based on the required balance between membrane anchoring, serum interaction, payload accessibility, and manufacturability. At this stage, we also identify the main technical risks, such as aggregation, over-lipidation, or active-site interference.
Chemistry is then matched to the project: amide coupling, thiol–maleimide reaction, click chemistry, or a building-block incorporation route. For liposome-related programs, we also compare direct coupling and post-insertion options when appropriate.
Because lipidated materials often behave differently from the parent molecule, purification is planned around amphiphilic properties rather than assumed from standard methods. We focus on removing free lipid, unconjugated payload, and partially modified species in a way that supports reliable downstream testing.
Final materials are examined for identity, purity, conjugation level, and—where relevant—particle attributes such as size and PDI. Comparative data can be generated for multiple lipid types, linker variants, or attachment positions to support design decisions rather than a single end-point readout.
Once an initial route is established, we support follow-on optimization, repeat synthesis, and workflow refinement for larger or more reproducible batches. This step is especially valuable when promising feasibility data must be translated into a more stable and better-characterized development package.
Why Clients Choose Our Lipid Conjugation Platform
We do not treat lipid conjugation as a one-format service. Our design logic can be adapted for oligonucleotides, peptides, proteins, small molecules, and lipid carrier surfaces so the chemistry matches the payload instead of forcing the payload into a generic workflow.

Lipidation can improve uptake or membrane interaction while creating new formulation problems. Our project planning emphasizes the trade-off between delivery benefit, aggregation risk, and functional retention so clients can compare options with clearer decision criteria.
We place equal weight on making the conjugate and proving what was made. That includes orthogonal review of identity, purity, free species, conjugation level, and particle-related attributes when working with liposome or PEG-lipid systems.
Our workflows are structured to support rapid proof-of-concept work first, then tighten stoichiometry control, purification, and comparative analytics as the project matures. This helps reduce rework when early hits need to be translated into more reliable material.
Application Areas of Lipid Conjugation
Oligonucleotide Delivery Optimization
- Lipid conjugation is widely used to improve the delivery behavior of siRNA, ASO, aptamers, and other nucleic acid formats.
- Appropriate lipid selection can enhance membrane interaction, tissue distribution characteristics, and compatibility with delivery systems.
- This application area is especially relevant when unconjugated oligonucleotides show limited uptake or insufficient delivery performance.
Liposome and Lipid Nanoparticle Surface Functionalization
- Lipid conjugation is commonly applied to attach peptides, proteins, oligonucleotides, or other ligands to liposome and lipid nanoparticle surfaces.
- It enables controlled ligand presentation for targeted delivery, surface engineering, and carrier customization.
- This direction is important for projects requiring reproducible surface loading and defined carrier functionality.
Peptide and Protein Lipidation
- Lipid conjugation is used to introduce hydrophobic anchors onto peptides and proteins for membrane association or altered solution behavior.
- This approach can support membrane-active construct design, surface presentation, and amphiphilic biomolecule development.
- It is particularly useful when the project requires stronger bilayer interaction or a defined lipid-mediated presentation format.
Membrane-Anchored Molecular Probe Development
- Lipid-conjugated probes are used in membrane biology, cell-surface studies, and localization-focused assay development.
- A lipid anchor can help position functional molecules at or near the membrane interface for more relevant biological evaluation.
- This application area includes imaging probes, reporter constructs, and membrane-associated assay tools.
Amphiphilic Biomolecule and Self-Assembly Research
- Lipid conjugation is an effective strategy for building amphiphilic molecules with controllable self-assembly behavior.
- It is used in projects involving supramolecular organization, nanostructure formation, and interface-driven biomolecular design.
- This direction is valuable when the goal is to combine biological function with lipid-mediated assembly properties.
Carrier-Compatible Functional Molecule Engineering
- Lipid conjugation is also used to adapt small molecules, ligands, and functional biomolecules for integration into lipid-based delivery or assay systems.
- It helps bridge the gap between an active molecule and the physical requirements of a carrier or membrane-associated platform.
- This application direction is relevant when a molecule must be reformatted for bilayer insertion, surface attachment, or lipid-assisted presentation.
Move Your Lipid Conjugation Project Forward with a Design-Focused Partner
Whether you are comparing lipid motifs for an oligonucleotide, building a lipidated peptide, attaching a protein to a reactive PEG-lipid, or improving ligand presentation on a liposomal system, we provide practical support from conjugation design through analytical confirmation.
Our team works with research groups to define workable chemistry, reduce avoidable formulation risk, and generate data packages that make next-step decisions easier. Contact our scientific team to discuss your lipid conjugation goals and build a project plan tailored to your molecule, carrier system, and development stage.
Frequently Asked Questions (FAQ)
Conjugated lipids enhance the solubility, stability, and cell membrane interaction of biomolecules, making them more effective for therapeutic or diagnostic applications.
Protein-lipid conjugates are biomolecules where lipids are attached to proteins to improve their membrane binding, cellular uptake, or therapeutic delivery efficiency.
Lipid-DNA conjugates are molecules where lipids are attached to DNA strands to enhance stability and delivery in gene therapy or nanotechnology applications.
Lipid conjugation improves drug solubility, protects active molecules from degradation, and enhances their targeted delivery across biological membranes.
Lipid conjugates are synthesized through chemical or enzymatic reactions, carefully designed to preserve the functionality of both the lipid and the attached biomolecule.

