Biotin Labeled Peptides
Site-Specific Peptide BiotinylationHigh-Purity Synthetic Peptide LabelsBuilt for ELISA, Pull-Down & Binding Assays
Advance assay development and peptide interaction studies with custom biotin labeled peptides designed for reliable immobilization, affinity capture, and sensitive detection. Biotinylated peptides combine defined synthetic peptide architecture with the strong biotin-streptavidin/avidin interaction, giving research teams a controlled way to anchor peptides onto coated plates, magnetic beads, membranes, microarrays, and biosensor surfaces for immunoassay, pull-down, epitope mapping, and proteomics workflows.
We support custom biotin labeled peptide synthesis for biotech, pharmaceutical, diagnostic, and academic research programs by integrating sequence review, labeling-site selection, linker engineering, purification, and analytical confirmation into one workflow. Projects can be tailored for N-terminal biotin, C-terminal Lys(Biotin), or internal side-chain biotinylation, with optional spacer designs such as Ahx or PEG-like linkers and compatible secondary modifications when required. Each construct is developed with downstream performance in mind so the final peptide is not only successfully labeled, but also fit for the assay it is meant to support.
What Problems Do Biotin Labeled Peptides Solve in Assay Development?
Many peptide-based workflows fail at the interface between chemistry and assay design: peptides adsorb poorly to assay surfaces, active motifs become masked after random immobilization, weak capture leads to low signal, and inconsistent presentation makes results difficult to reproduce. Biotin labeled peptides address these issues by providing a defined attachment point that enables stable capture on streptavidin or avidin systems while preserving control over peptide orientation, spacing, and accessibility. This is especially valuable for peptide ELISA, pull-down experiments, binding kinetics studies, peptide microarrays, overlapping peptide screening, and affinity enrichment workflows where assay performance depends on how the peptide is presented rather than on sequence alone.
 Fig.1 The structures of the designed biotinylated peptides. (Matsumura et al., 2005)
Fig.1 The structures of the designed biotinylated peptides. (Matsumura et al., 2005)
Key Challenges in Custom Biotinylated Peptide Projects
A peptide may synthesize cleanly yet fail in the assay if biotin is attached at a terminus or residue that participates in target recognition. We review sequence context, known binding residues, and intended assay geometry to recommend a labeling site that minimizes the risk of masking the biologically relevant region.
Direct biotin attachment is not always enough. If the peptide sits too close to the streptavidin-coated surface, binding partners may not access the active sequence efficiently. Spacer-assisted formats such as Ahx or PEG-like linkers are often needed to improve presentation and assay sensitivity.
Hydrophobic, aggregation-prone, highly basic, or multi-modified peptides can be harder to synthesize, purify, and reconstitute after labeling. Sequence-aware design and the right linker architecture help reduce handling issues and improve the likelihood of obtaining an assay-usable product.
In pull-down, ELISA, and biosensor work, the difference between a cleanly labeled peptide and a partially modified or impure material can translate directly into background, weak response, or poor reproducibility. We emphasize HPLC and LC-MS-based release data so teams can move forward with clearer confidence in identity, purity, and labeling integrity.
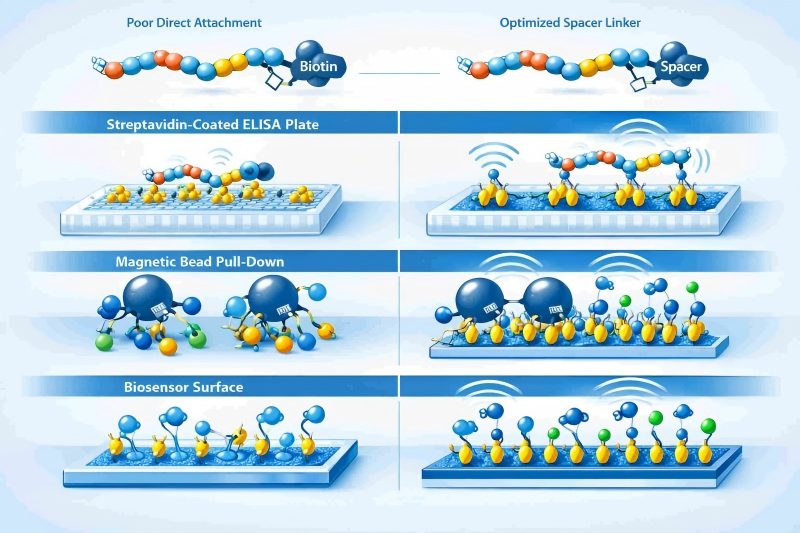 Spacer-optimized biotin labeled peptides help improve peptide presentation and assay consistency across streptavidin-based research platforms.
Spacer-optimized biotin labeled peptides help improve peptide presentation and assay consistency across streptavidin-based research platforms.
Our Biotin Labeled Peptide Services
We provide assay-oriented custom biotin labeled peptide development for research teams that need more than a standard modification add-on. In addition to broader Biotinylation and Peptide Conjugation Services, this service is focused specifically on peptide constructs where labeling position, linker choice, purification strategy, and analytical release all need to align with downstream performance.
 Biotinylation Strategy Design & Sequence Review
Biotinylation Strategy Design & Sequence Review
Capabilities include:
- Evaluation of peptide sequence, active motif location, and terminal accessibility
- Selection of N-terminal, C-terminal, or internal side-chain biotinylation strategy
- Spacer recommendation based on assay surface, capture format, and steric requirements
- Design input for ELISA, pull-down, peptide array, SPR, and BLI workflows
- Planning of labeled and unlabeled control peptides when comparative studies are needed
- Guidance on purity targets, quantity, and scale according to project stage
Best suited for:
New assay setup, low-signal troubleshooting, and projects where peptide function must be preserved after biotin incorporation
 N-Terminal Biotin Labeled Peptide Synthesis
N-Terminal Biotin Labeled Peptide Synthesis
Capabilities include:
- Solid-phase synthesis of peptides bearing N-terminal biotin or biotin-linker motifs
- Consistent construct design for single peptides or larger screening panels
- Optional C-terminal amidation or other terminal adjustments where sequence context requires it
- Purification strategy aligned to desired analytical quality
- HPLC purification and LC-MS confirmation of final labeled products
- Delivery formats tailored to research and assay development use
Typical applications:
Peptide ELISA, plate immobilization, peptide arrays, overlapping peptide panels, and general streptavidin-based detection workflows
 C-Terminal & Internal Lys(Biotin) Peptide Design
C-Terminal & Internal Lys(Biotin) Peptide Design
Capabilities include:
- C-terminal Lys(Biotin) incorporation when the N-terminus should remain unmodified
- Internal lysine-side-chain biotinylation for site-controlled presentation
- Orthogonal design approaches to place the label away from the functional epitope
- Linker insertion before Lys(Biotin) when additional separation is beneficial
- Sequence-aware design for receptor binding and competitive interaction studies
- Analytical confirmation that the intended modified species is obtained
Typical applications:
Pull-down assays, receptor-ligand studies, affinity enrichment, and projects where terminal freedom is important for activity
 Spacer-Optimized Biotinylated Peptides
Spacer-Optimized Biotinylated Peptides
Capabilities include:
- Direct biotin, Biotin-Ahx, and PEG-like spacer options for different surface-access requirements
- Design to reduce steric hindrance between peptide and streptavidin-coated supports
- Hydrophilic linker selection to improve solubility and handling of difficult sequences
- Comparative design of multiple spacer formats when assay fit is uncertain
- Optimization support for sensor immobilization and bead-based capture
- Coordination of dual-labeled concepts with Fluorescence Labeling of Peptides when orthogonal readouts are needed
Focus areas:
Improved peptide accessibility, better assay signal, and more reproducible target engagement on coated surfaces
 Biotinylated Peptide Panels & Multi-Modified Constructs
Biotinylated Peptide Panels & Multi-Modified Constructs
Capabilities include:
- Overlapping peptide sets, truncation series, and sequence-variant panels with consistent biotin formatting
- Biotinylated constructs combined with amidation, acetylation, phosphorylation, cyclization, or disulfide features where relevant
- Matched unlabeled or alternative-site controls for structure-function comparison
- Design support for epitope mapping and interaction screening campaigns
- Flexible incorporation strategies using appropriate modified residues and building blocks
- Related raw-material support can be aligned with Biotinylated Amino Acid Services for specialized projects
Typical applications:
Epitope mapping, peptide microarrays, motif validation, protein interaction screening, and assay panel development
 Purification, Analytical Characterization & Assay-Ready Release
Purification, Analytical Characterization & Assay-Ready Release
Capabilities include:
- Preparative and analytical HPLC workflows suited to labeled peptide purification
- LC-MS identity confirmation of the target biotinylated construct
- Review of major truncated, deleted, or incompletely modified species where detectable
- Purity targets aligned to assay intent, with many binding and immunoassay programs requesting high-purity material
- Batch-specific documentation for repeat ordering and method continuity
- Lyophilized delivery with sequence-aware handling and reconstitution guidance
Deliverables:
HPLC chromatograms, LC-MS data, purity summary, and project-specific release information suitable for internal research documentation
Recommended Biotinylation Formats by Application
The most effective biotin labeled peptide is not always the simplest one. Attachment site, spacer choice, and peptide architecture should be matched to the assay format, the location of the active motif, and the handling characteristics of the sequence. The table below summarizes common design choices used to improve fit with real research workflows.
| Application Goal | Common Labeling Format | Preferred Spacer Strategy | Why It Is Often Chosen | What Clients Should Clarify Up Front |
| Peptide ELISA / Plate Coating | N-terminal biotinylation when the N-terminus is not part of the epitope | Ahx or hydrophilic spacer frequently useful | Supports stable immobilization and more consistent plate presentation | Antibody target region, plate type, desired purity, and whether an unlabeled control is needed |
| Pull-Down / Affinity Capture | Terminal or internal site-specific biotin away from the interaction motif | Spacer often recommended to improve bead-accessible presentation | Helps preserve binding while enabling efficient streptavidin-based capture | Sample source, wash stringency, control design, and downstream readout such as WB or MS |
| SPR / BLI Surface Immobilization | Terminus selected to orient the binding face away from the sensor surface | Short or hydrophilic linker depending on kinetics format | Improves interpretability of affinity and kinetic data by controlling presentation | Sensor platform, regeneration conditions, and known critical residues |
| Epitope Mapping Panels | Consistent N-terminal biotin across the full peptide set | Same linker format for all panel members | Reduces variation caused by inconsistent immobilization across overlapping peptides | Peptide list, overlap length, terminal capping needs, and panel scale |
| Receptor / Competitive Binding Studies | C-terminal or side-chain biotinylation when the N-terminus must remain free | Spacer selected according to steric sensitivity of the target system | Maintains access to the pharmacophore or receptor-recognition segment | Known active motif, binding model, and whether parallel alternative-site constructs are desired |
| Difficult Hydrophobic Sequences | Labeling site chosen with handling and purification behavior in mind | PEG-like linker often considered to improve accessibility and solubility | Can reduce aggregation risk and support cleaner assay performance | Solvent restrictions, reconstitution preference, and acceptable formulation conditions |
Biotin Incorporation Strategies & Synthesis Considerations
Biotin can be introduced into peptides in multiple ways, and each approach affects accessibility, synthesis complexity, and downstream assay behavior differently. The following overview highlights practical options commonly considered in custom biotin labeled peptide design.
| Incorporation Strategy | Technical Approach | Where It Is Most Useful | Key Considerations |
| Direct N-Terminal Biotin | Biotin introduced at the peptide N-terminus during solid-phase synthesis | Standard screening peptides and straightforward immobilization formats | Efficient format, but not ideal when the N-terminus is part of the active binding region |
| N-Terminal Biotin-Ahx | Biotin separated from the peptide by an aminohexanoic acid spacer | ELISA, pull-down, and bead-based assays that benefit from extra distance to the surface | Common choice when direct attachment causes steric hindrance or weak signal |
| N-Terminal Biotin-PEG Format | Biotin linked to the peptide through a short PEG-like hydrophilic spacer | Difficult or hydrophobic sequences and assays requiring improved accessibility | Can improve solubility and presentation, but the best linker still depends on the sequence and assay setup |
| C-Terminal Lys(Biotin) | Additional lysine at the C-terminus carries the biotin on the side chain | Cases where the native N-terminus should remain unmodified | Useful for preserving N-terminal function while retaining a defined terminal attachment point |
| Internal Lys(Biotin) / Side-Chain Labeling | Biotin introduced on a designated lysine residue within the sequence | Site-directed projects requiring controlled placement away from terminal regions | Requires careful residue selection so the label does not disrupt folding, recognition, or charge balance |
| Biotin Plus Secondary Modification | Biotin combined with other peptide features such as phosphorylation, amidation, cyclization, or fluorescent tags | Advanced structure-function studies and assay-specific probe design | Demands tighter synthesis planning and analytical review because each added modification changes project risk |
Analytical Characterization & Release Package for Biotinylated Peptides
For biotin labeled peptides, analytical confirmation is not just a documentation step. It is a practical control point that helps determine whether the peptide entering the assay is the one originally designed. Our characterization framework is built to support research programs that require confidence in identity, purity, modification integrity, and repeatability across repeat orders or panel builds.
| Analytical Category | Methodology | Why It Matters | Typical Data Delivered |
| Identity Confirmation | LC-MS molecular weight analysis | Confirms that the intended biotinylated peptide species has been obtained | Mass spectrum and calculated/found molecular weight summary |
| Purity Assessment | Analytical HPLC | Helps evaluate the level of full-length target material relative to detectable impurities | Chromatogram and purity percentage summary |
| Label Incorporation Review | HPLC and LC-MS correlation | Provides evidence that the peptide carries the expected biotin modification rather than an unlabeled or incompletely modified species | Analytical interpretation of labeled target peak and major related species |
| Secondary Modification Check | LC-MS and project-specific chromatographic review | Important for constructs that include phosphorylation, cyclization, amidation, or other additional features | Modification-integrity confirmation aligned to project scope |
| Repeat-Order Comparability | Cross-lot analytical comparison when applicable | Supports consistency in ongoing assay programs or larger panel expansions | Batch comparison notes and repeat-order analytical continuity |
| Release Documentation | Structured project reporting | Gives research teams traceable data for internal records and downstream assay planning | COA-style summary, chromatograms, mass data, and handling information |
Workflow for Custom Biotin Labeled Peptide Development
We start by reviewing the peptide sequence, the known or suspected functional region, the assay platform, and the role the biotinylated construct must play. This helps identify whether the main risk is motif masking, poor capture, low solubility, or insufficient analytical confidence.
Based on project goals, we recommend the most suitable biotinylation site and whether a direct label, Ahx spacer, or PEG-like linker is more appropriate. When needed, we can also suggest alternative-site constructs or unlabeled controls to improve data interpretation.
The peptide is synthesized with the selected biotin format using a route compatible with the sequence complexity and any additional requested modifications. Attention is given to difficult residues, side-reaction risk, and label integration so the construct remains aligned with the original design intent.
Purification conditions are selected to obtain the level of material quality appropriate for the requested use. This stage is particularly important for biotinylated and multi-modified peptides, where truncated or incompletely modified species can affect downstream assay behavior.
Analytical testing is performed to confirm molecular identity and assess purity. For repeat projects or assay-critical constructs, the resulting release data also supports cross-project consistency and better decision-making before the peptide enters experimental workflows.
Peptides are delivered with the relevant analytical package and handling information. We can continue to support follow-on work such as sequence variants, alternate linker designs, larger panels, or related capture-system projects as assay development evolves.
Why Research Teams Choose Our Biotinylated Peptide Platform
We do not treat biotinylation as a generic end modification. Label placement and linker selection are aligned to the real experimental context so the final construct is better suited to plate capture, bead pull-down, biosensor loading, or screening workflows.

Synthetic peptide design allows biotin to be placed precisely at the N-terminus, C-terminus, or an intended lysine side chain. This supports more homogeneous materials than random labeling approaches and makes comparative design work more meaningful.
Hydrophobic motifs, sterically sensitive binders, multi-modified peptides, and panel-based projects often need more thoughtful design than standard catalog products provide. We build around those constraints from the start rather than after the assay fails.
Our workflow combines synthesis, purification, and analytical confirmation so clients receive data-backed peptide materials rather than modification claims alone. This is especially important for teams that need cleaner handoff into assay development and repeat ordering.
Common Applications of Biotin Labeled Peptides
Peptide ELISA & Immunoassay Development
- Immobilization of peptide antigens or probes on streptavidin-coated plates.
- Epitope-specific antibody screening and assay optimization.
- More consistent surface presentation than non-specific adsorption-based coating.
Pull-Down & Affinity Enrichment
- Capture of peptide-binding proteins using streptavidin beads or related affinity supports.
- Enrichment workflows for Western blot or mass spectrometry follow-up.
- Useful for ligand discovery, partner confirmation, and interaction validation.
SPR, BLI & Biosensor Surface Loading
- Controlled immobilization of peptides for real-time binding analysis.
- Better orientation control for kinetic and affinity measurements.
- Compatible with projects that need comparative linker or site-position studies.
Epitope Mapping & Overlapping Peptide Studies
- Uniformly formatted peptide panels for antibody or protein-binding region mapping.
- Support for overlapping libraries, truncation series, and sequence variants.
- Well suited to plate-based and array-based screening formats.
Peptide Microarrays & High-Throughput Screening
- Surface-ready peptides for parallel binding and specificity studies.
- Consistent biotin presentation across larger peptide sets.
- Useful for screening motifs, interaction hotspots, and sequence-function relationships.
Proteomics & Target Identification Workflows
- Affinity capture of interacting partners from complex biological samples.
- Support for downstream SDS-PAGE, immunoblotting, and MS analysis.
- Practical when a defined peptide bait is preferred over larger protein constructs.
Start Your Biotin Labeled Peptide Project with a Fit-for-Assay Design
Whether you need a single biotinylated peptide for a peptide ELISA, a spacer-optimized bait for pull-down studies, or a larger overlapping panel for epitope mapping, we provide design-aware synthesis and analytical support tailored to the way the material will actually be used.
Our team works with research groups to refine labeling position, improve accessibility on streptavidin-based platforms, and deliver cleaner peptide reagents for downstream binding and detection studies. Where relevant, projects can also be coordinated with related Streptavidin Conjugation or Biotin-Streptavidin Conjugation support.Contact our scientific team to discuss your biotin labeled peptide sequence, assay format, and preferred labeling strategy.
Frequently Asked Questions (FAQ)
Biotinylation efficiency can be optimized by selecting the appropriate biotin reagent, adjusting reaction conditions such as pH, temperature, and reaction time. Using high-quality peptides and ensuring minimal steric hindrance at the labeling site also improves efficiency.
Yes, biotin-labeled peptides are ideal for cell surface binding assays, as the biotin-streptavidin interaction allows for strong and specific binding, facilitating easy detection and quantification of peptide interactions with cell surface receptors.
To minimize non-specific binding, use blocking agents like BSA or non-fat dry milk during your experiments. Additionally, optimize peptide concentration and ensure proper buffer conditions to reduce background noise during assays.

