Fluorescence Labeling of Nucleic Acids
Site-Controlled Fluorophore AttachmentCustom DNA & RNA Probe EngineeringFrom Single Labels to Dual-Labeled FRET Designs
Advance nucleic acid assay development, imaging workflows, and probe engineering with fluorescence labeling of nucleic acids tailored to the needs of research teams, assay developers, and bioconjugation programs. Fluorescently labeled DNA, RNA, and oligonucleotide constructs are widely used for hybridization-based detection, qPCR probe design, FISH, uptake tracking, intracellular localization studies, and structure-function analysis because they enable sensitive, multiplexable, non-radioactive readouts across diverse experimental platforms.
We support custom labeling of synthetic oligonucleotides and selected DNA/RNA constructs using 5′ labeling, 3′ labeling, internal labeling, dual-labeled probe formats, and post-synthetic or synthesis-integrated conjugation strategies. Projects can be configured around dye family, labeling position, spacer/linker design, purification level, and analytical release criteria. For adjacent needs, we also support broader fluorescence labeling, fluorescence labeling of oligonucleotides, and oligonucleotide bioconjugation workflows. When a non-fluorescent capture format is a better fit for the downstream assay, related options such as biotin-labeled nucleic acids can also be considered.
What Practical Problems Does Fluorescence Labeling of Nucleic Acids Solve?
Fluorescence labeling is not just about attaching a dye to DNA or RNA. It is a problem-solving step that helps research teams overcome weak signal, poor signal-to-background ratio, cross-channel interference, and probe formats that are difficult to interpret in real assays. Many nucleic acid projects fail because the fluorophore is installed at the wrong position, the dye family does not match the instrument optics, the label perturbs hybridization or RNA performance, or the final material still contains free dye and unlabeled species. A well-designed fluorescence labeling workflow addresses these issues by aligning sequence context, labeling position, spacer strategy, fluorophore selection, purification method, and QC expectations. The result is a labeled construct that is easier to detect, easier to validate, and more reliable in applications such as qPCR, FISH, imaging, duplex tracking, uptake studies, and biosensor development.
 Fig.1 Fluorescent labeling of nucleic acids. (Rombouts, K., 2016)
Fig.1 Fluorescent labeling of nucleic acids. (Rombouts, K., 2016)
Key Challenges in Fluorescence Labeling of Nucleic Acids
Brightness on paper does not always translate into clean assay performance. Fluorophore quenching, photobleaching, incomplete purification, or mismatch between dye choice and instrument channel can produce low apparent signal or high background. We help align dye family, label density, and purification strategy with the intended readout system.
A fluorescent tag placed too close to a critical hybridization region, enzymatic recognition site, or structured RNA domain can reduce assay performance. We evaluate whether 5′, 3′, internal, or dual-label formats are more appropriate for the sequence architecture and downstream use.
Multi-color assays require careful control of spectral overlap, quencher compatibility, and donor-acceptor spacing. We support rational probe-format selection for dual-labeled hydrolysis probes, molecular beacons, and FRET-oriented constructs to reduce rework during assay optimization.
Free dye, partially labeled species, truncated sequences, and duplex assembly variability can compromise interpretation. Our workflows are designed around appropriate purification and analytical confirmation so teams receive labeled material with clearer identity, purity, and labeling-status assessment before deployment.
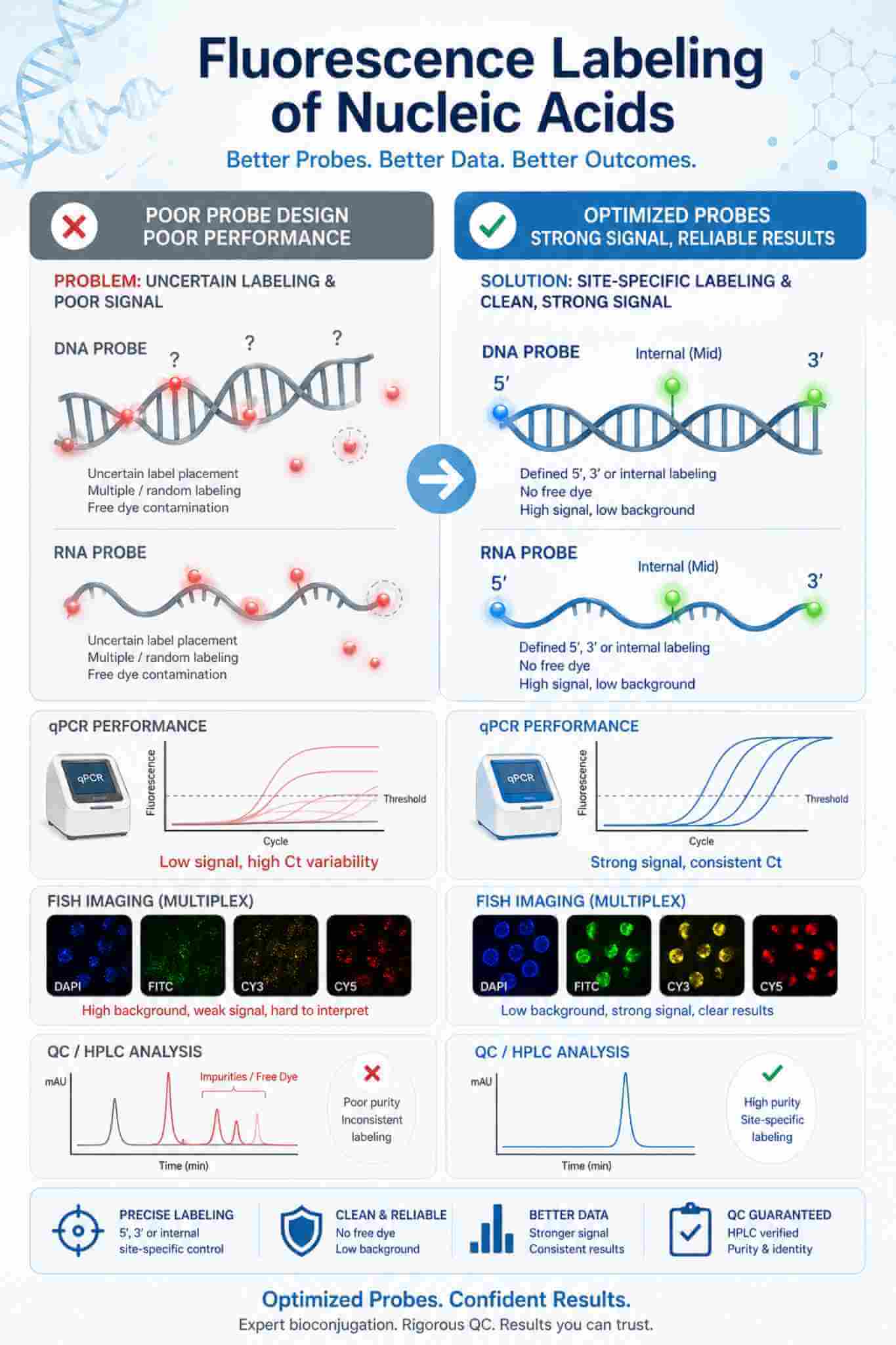 Rational dye selection, site-controlled labeling, and proper purification help convert fluorescent nucleic acids from inconsistent reagents into reliable tools for detection, imaging, and probe-based assays.
Rational dye selection, site-controlled labeling, and proper purification help convert fluorescent nucleic acids from inconsistent reagents into reliable tools for detection, imaging, and probe-based assays.
Our Fluorescence Labeling Services for DNA, RNA, and Oligonucleotide Projects
We provide custom fluorescence labeling services for nucleic acid projects ranging from single-labeled oligonucleotides to dual-labeled probes and selected longer DNA/RNA constructs. Each program is planned around application fit, sequence context, labeling chemistry, purification strategy, and analytical verification so the final material is matched to real assay needs rather than treated as a generic modification order.
 Fluorescent Oligonucleotide Synthesis & Site-Specific Labeling
Fluorescent Oligonucleotide Synthesis & Site-Specific Labeling
Capabilities include:
- 5′, 3′, and internal fluorescent labeling for custom DNA and RNA oligonucleotides
- Selection of dye family and attachment mode based on assay format and detection channel
- Spacer and linker considerations to reduce steric interference and improve accessibility
- Single-label and multi-position designs where sequence context allows
- Purification by HPLC or PAGE as appropriate for sequence length and modification profile
- Identity and labeling confirmation by LC-MS and spectroscopic assessment for suitable constructs
- Support for DNA probes, RNA probes, adapters, primers, and capture oligos
Typical applications:
Hybridization probes, labeled primers, imaging oligos, sequencing-supporting oligos, and general fluorescence-based nucleic acid assays
 Dual-Labeled Probes, Molecular Beacons & FRET Constructs
Dual-Labeled Probes, Molecular Beacons & FRET Constructs
Capabilities include:
- Reporter-quencher probe design for hydrolysis probes and related qPCR formats
- Molecular beacon and stem-loop style fluorescent oligonucleotide preparation
- Donor-acceptor pair selection for FRET-oriented nucleic acid studies
- Sequence and spacing review to improve quenching efficiency and target response
- Multiplex-oriented dye set planning to reduce spectral crosstalk
- Purification and QC adapted to dual-labeled constructs
- Design support for SNP detection, probe-based amplification, and conformational readouts
Typical applications:
qPCR probe development, molecular beacon assays, hybridization-triggered fluorescence, and nucleic acid FRET studies
 Fluorescent siRNA and Functional RNA Labeling
Fluorescent siRNA and Functional RNA Labeling
Capabilities include:
- Fluorescent labeling of siRNA, short RNA oligos, and related functional RNA constructs
- Terminal or strand-selective labeling strategies chosen to reduce performance disruption
- Labeling plans for uptake, trafficking, transfection monitoring, and localization studies
- Handling logic suited to RNA sensitivity and downstream duplex requirements
- Optional control-material planning for comparative assay interpretation
- Analytical verification of labeled RNA identity and purity for suitable project formats
Focus areas:
Cellular uptake studies, intracellular distribution analysis, transfection optimization, and fluorescence-based RNA workflow development
 Longer DNA/RNA Probe Labeling & Analytical QC Support
Longer DNA/RNA Probe Labeling & Analytical QC Support
Capabilities include:
- Post-synthetic labeling strategies for modified nucleic acids where direct synthesis is not the best route
- Enzymatic incorporation approaches for selected longer DNA or RNA probe formats
- Review of labeling density requirements versus hybridization performance
- Cleanup and purification planning for removal of free dye and under-labeled material
- Analytical characterization using chromatography, gel-based methods, UV-Vis, and related techniques as appropriate
- Comparative planning when fluorescent tags are being evaluated against alternative formats such as biotin-labeled nucleic acids
Deliverables:
Labeled nucleic acid material supported by project-relevant purity assessment, labeling-status evaluation, and technical reporting for research use
Choosing the Right Labeling Format for Nucleic Acid Fluorescence Projects
Successful fluorescence labeling starts with choosing a format that matches how the nucleic acid will actually be used. Position, label count, and installation route can change hybridization behavior, assay background, and analytical complexity. The table below summarizes common design options and when each is most useful.
| Labeling Format | Best Suited For | Main Advantages | Key Design Considerations | Typical Use Cases |
| 5′ End Labeling | Primers, probes, adapters, short DNA/RNA oligos | Straightforward design, broad compatibility, usually lower risk of hybridization disruption | Confirm the 5′ terminus is not critical for enzymatic handling or interaction with other components | qPCR probes, imaging probes, labeled primers, uptake tracers |
| 3′ End Labeling | Oligos requiring a terminal label away from the main recognition region | Useful when the 5′ end must remain unmodified or when probe geometry favors a 3′ fluorophore | Check compatibility with polymerase, ligase, extension, or nuclease-related workflow requirements | Hybridization probes, protected terminal formats, certain tracking oligos |
| Internal Labeling | Signal-tuned probes, structure studies, specialized assay formats | Can improve spatial placement of the reporter and enable more advanced probe behavior | Internal modifications may affect duplex stability, sequence composition, or local structure | FRET studies, structure-sensitive probes, specialized hybridization assays |
| Dual-Labeled Probe | Reporter-quencher systems and multiplex assay development | Enables turn-on or cleavage-based signal generation with tighter background control | Requires matched reporter/quencher pairing, spacing logic, and more stringent QC | Hydrolysis probes, molecular beacons, SNP assays, real-time amplification |
| Post-Synthetic Chemical Labeling | Amine- or thiol-modified oligos and custom follow-on conjugation workflows | Flexible dye choice after nucleic acid synthesis and useful for custom project adaptation | Requires efficient coupling and cleanup to remove free dye and under-labeled material | Custom probe screening, late-stage dye substitution, special fluorophore installs |
| Enzymatic Incorporation | Selected longer DNA/RNA probes and amplified labeling workflows | Useful when multiple dye incorporations or longer templates are needed | Labeling density and incorporation pattern must be balanced against target performance | FISH probes, amplified DNA probes, labeled templates for hybridization studies |
Fluorophore Selection Considerations for DNA and RNA Labeling
Dye selection affects much more than color. Brightness, photostability, pH behavior, compatibility with filters and detectors, quencher pairing, and multiplex separation all influence whether a fluorescent nucleic acid performs well in practice. The table below outlines common fluorophore classes and the selection logic typically considered during project design.
| Dye Class / Region | Common Examples | Typical Strengths | Common Uses | Selection Notes |
| Green Channel | Fluorescein / FAM-like dyes | High familiarity, broad instrument compatibility, strong signal in many standard platforms | General oligo labeling, probe development, routine fluorescence readouts | Often a practical starting point, but pH sensitivity and photostability should still be considered |
| Yellow-Green Channel | HEX / JOE-like dyes | Useful for multiplex separation from green-channel reporters | Multi-color qPCR and parallel probe detection | Good when the platform already uses a green reporter and an additional channel is needed |
| Orange Channel | TAMRA / Cy3-like dyes | Strong visibility for imaging-oriented and probe-based workflows | Cell-associated tracking, imaging probes, multiplex assay sets | Evaluate spectral overlap carefully when combined with green or red fluorophores |
| Red / Far-Red Channel | Cy5 / Alexa 647-like dyes | Valuable for low-background imaging and multi-color designs requiring spectral separation | FISH, cell imaging, tracking experiments, higher-order multiplex panels | Often chosen when reduced background or deeper channel separation is important |
| Quencher Pairing | BHQ-, Dabcyl-, and matched quencher systems | Reduces background signal in dual-labeled probe formats | Hydrolysis probes, molecular beacons, turn-on hybridization assays | Quencher choice must be matched to the reporter emission profile and probe architecture |
Analytical Characterization Framework for Fluorescently Labeled Nucleic Acids
Fluorescent nucleic acid projects need more than sequence confirmation. The analytical package should distinguish labeled product from free dye, truncated material, unlabeled sequence, and partially modified species wherever the construct type allows. Our QC framework is selected according to nucleic acid format, sequence length, label complexity, and intended use.
| Analytical Category | What It Confirms | Typical Methods | Project Value |
| Identity Verification | Confirms the expected labeled nucleic acid species for suitable construct types | LC-MS, mass-based confirmation, sequence-correlated analytical review | Reduces uncertainty before assay deployment and helps distinguish correct product from synthesis by-products |
| Purity Assessment | Evaluates full-length product relative to truncated and process-related impurities | HPLC, UPLC, PAGE, capillary or gel-based methods as appropriate | Improves interpretability in sensitive fluorescence readouts and hybridization assays |
| Free Dye / Under-Labeled Species Control | Determines whether unreacted fluorophore or incompletely labeled material remains | Chromatographic separation, gel analysis, spectroscopic review | Helps reduce background signal and unexpected assay behavior |
| Dye-to-Oligo Spectral Review | Confirms absorbance behavior and labeling presence from a fluorescence/UV perspective | UV-Vis and fluorescence-based measurements | Supports downstream concentration setting, probe handling, and comparative assay normalization |
| Duplex / Assembly Check | Reviews whether duplexed or paired probe constructs assemble as intended | Gel-shift analysis, hybridization check, project-specific functional screening | Useful for siRNA, beacon formats, and other multi-component nucleic acid systems |
| Application-Oriented Release Review | Aligns analytical output with how the material will actually be used | Combined review of purity, labeling status, sequence, and assay-fit criteria | Helps research teams move from synthesis to experiment with fewer avoidable redesign cycles |
Workflow for Custom Fluorescence Labeling of Nucleic Acids
We start by reviewing nucleic acid type, sequence length, structure risk, intended assay, instrument channel, and whether the construct will be used as a primer, probe, duplex, or imaging reagent. This step helps prevent avoidable label-placement mistakes before synthesis begins.
Based on the project goal, we recommend 5′, 3′, internal, or dual-label designs together with fluorophore and quencher combinations that match detection hardware, multiplex requirements, and sequence constraints.
The construct is prepared using the most suitable route for the material type, including synthesis-integrated labeling, post-synthetic modification of reactive handles, or selected enzymatic incorporation strategies for longer probe formats.
We apply purification methods appropriate to the sequence and modification profile to reduce free fluorophore, truncated material, and incompletely labeled species that can otherwise distort fluorescence readouts.
Labeled products undergo identity, purity, and labeling-status review using analytical tools selected for the construct class. This stage is essential for distinguishing a visually fluorescent sample from an analytically reliable one.
Final materials are delivered with project-relevant documentation and handling recommendations so teams can move directly into assay optimization, imaging, or probe validation with clearer expectations for storage, light protection, and experimental setup.
Why Choose Our Nucleic Acid Fluorescence Labeling Service
We do not treat fluorescent labeling as a simple afterthought. Label placement is evaluated against hybridization regions, duplex architecture, assay format, and potential steric effects so the final construct is more likely to work in the intended experiment.

Our scope spans single-labeled DNA/RNA oligos, dual-labeled probe systems, fluorescent siRNA, and selected longer nucleic acid labeling strategies, allowing project teams to stay within one technical workflow as needs evolve.
Fluorescent nucleic acids often require different analytical thinking than unmodified sequences. We select purification and characterization methods according to label complexity, sequence length, and downstream sensitivity rather than relying on one default release model.
Whether the priority is channel separation, quencher pairing, intracellular visualization, or cleaner interpretation in probe-based assays, we can adapt the labeling strategy to the actual readout environment and project decision points.
Applications of Fluorescence-Labeled Nucleic Acids
qPCR and Hybridization Probe Development
- Reporter-quencher probes for fluorescence-based amplification workflows.
- Sequence-specific probes for endpoint or real-time hybridization assays.
- Multiplex probe sets requiring channel separation and spectral planning.
FISH and Spatial Nucleic Acid Imaging
- Fluorescent DNA or RNA probes for localization and visualization studies.
- Probe sets designed for cell-based imaging and fixed-sample workflows.
- Color-channel planning for multi-target imaging panels.
Uptake, Transfection, and Intracellular Tracking
- Fluorescent siRNA and RNA constructs for monitoring delivery behavior.
- Tracking of nucleic acid localization and trafficking in cell studies.
- Comparative use of labeled control oligos in optimization workflows.
Biosensors, FRET, and Structure-Function Studies
- Donor-acceptor labeled nucleic acids for conformational and interaction studies.
- Fluorogenic or quenched probe formats for target-responsive readouts.
- Custom labeled constructs for assay development in analytical and biochemical research.
Case Study
Case Study 1
Chemical selective modification reactions are widely used in the labeling of biomolecules, including copper-catalyzed or tension-promoted azide-alkyne cycloaddition (CuAAC/SPAAC) and Diels-Alder (IEDDA) reaction with anti-electron demand. Although these reactions can be effectively labeled in vitro, the labeling of nucleic acids in vivo is still limited to single-stranded RNA or requires membrane-permeable or phototoxic methods to efficiently label chromatin-encapsulated double-stranded DNA (dsDNA). In this study, the author designed a new imaging probe (PINK) for nucleoside olefin groups. PINK consists of fluorescent embedded reagent and tetrazine group. Tetrazine group is a bioorthogonal group and a fluorescence quenching group, and the fluorescence embedded reagent partially enables it to reversibly interact with double-stranded DNA until it is connected with olefin-containing nucleotides. The combination of the two can label double-stranded DNA very quickly and fluorescently.
 Fig. 2 Schematic diagram of PINK probe. (Loehr, M. O., 2022)
Fig. 2 Schematic diagram of PINK probe. (Loehr, M. O., 2022)
Case Study 2
Marcus Wilhelmsson et al. synthesized a tricyclic cytosine analog with fluorescence properties, called tCO. The excitation wavelength was 369nm, and the strongest fluorescence wavelength was 457nm. It can be doped into the target RNA molecule by terminal deoxynucleotidyl transferase (TdT) or RNA polymerase to realize internal fluorescence labeling. Different from other fluorescent markers, tCOTP is more similar to natural CTP, which can be correctly recognized by RNA polymerase or deoxynucleotidyl transferase and the translation machine in the cell, with little interference to the translation process, and can be correctly folded and located in the right position in the cell. More importantly, for the first time, they realized the global living cell imaging of mRNA molecules labeled with fluorescent base analogues from cell uptake to protein expression, which will bring great help to the mechanism of cell uptake and intracellular escape. It is expected that this convenient, universal and minimally interfering fluorescent labeling method can be applied to the development of nucleic acid delivery systems and promote the upgrading of RNA drugs.
 Fig. 3 Translation of tCO-labeled mRNA molecules in human cells by chemical transfection. (Baladi, T., 2021)
Fig. 3 Translation of tCO-labeled mRNA molecules in human cells by chemical transfection. (Baladi, T., 2021)
Discuss Your Fluorescence Labeling Strategy for DNA, RNA, and Oligonucleotide Projects
Whether you need a single fluorescently labeled oligonucleotide, a dual-labeled probe set, a fluorescent siRNA construct, or a more customized DNA/RNA labeling plan, we can help define a strategy that fits the real demands of your assay or imaging workflow.
Our team works from sequence context, probe format, dye compatibility, purification requirements, and analytical expectations to reduce redesign cycles and improve the usefulness of the final labeled material. For teams evaluating broader design options, we can also help connect the project with related resources such as fluorescent labeling technology and adjacent nucleic acid conjugation approaches. Contact our scientific team to discuss your fluorescence labeling of nucleic acids project and request a tailored technical plan.
Frequently Asked Questions (FAQ)
Fluorescence labeling enables researchers to track and study nucleic acids in vitro and in vivo. It is valuable for applications such as in situ hybridization, fluorescence in situ hybridization (FISH), DNA sequencing, real-time PCR, and various cellular imaging techniques.
Nucleic acids can be labeled by directly incorporating fluorescently modified nucleotides during synthesis (e.g., in PCR or sequencing), or by post-synthetic labeling methods where a fluorescent dye or probe is attached to the nucleic acid molecule using chemical conjugation techniques.
Some popular fluorescent dyes used for nucleic acid labeling include fluorescein (FITC), Cy3, Cy5, Texas Red, SYBR Green, and various Alexa Fluor dyes. These dyes emit light at specific wavelengths when excited by a light source.
Fluorescence labeling generally does not significantly alter the properties or functions of nucleic acids if performed correctly. However, bulky or inappropriate labeling may interfere with hybridization or enzymatic reactions.
Labeled nucleic acids can be visualized using fluorescence microscopy, fluorescence plate readers, or imaging systems that detect specific emission wavelengths corresponding to the fluorescent dye used.
Compatibility with downstream applications
Sensitivity and specificity of the labeling method
Impact on nucleic acid function
Stability and photostability of the fluorescent label
Cost and ease of use
Limitations include potential interference with nucleic acid function, background fluorescence, photobleaching (loss of fluorescence over time), and the need for optimization based on specific experimental requirements.