Aptamer Conjugation Services
Programmable Target RecognitionFlexible Linker & Payload DesignCustom Aptamer Conjugation for Delivery, Assays & Nanoplatforms
Accelerate targeted research and advanced molecular system design with custom aptamer conjugation services for biotech, pharmaceutical, diagnostics, and platform development teams. Aptamers are sequence-defined DNA or RNA ligands that can be conjugated to drugs, fluorophores, proteins, nanoparticles, liposomes, beads, and complementary oligonucleotides to create highly selective tools for delivery, binding analysis, enrichment, imaging, and biosensing.
We support aptamer conjugation projects from sequence and structure review through conjugation route selection, linker design, purification, and analytical confirmation. Whether your goal is to preserve aptamer binding after payload attachment, control surface density on a carrier, or generate a robust labeled probe for downstream assays, our workflows are tailored to the chemistry, application, and performance requirements of each project.
What Problems Does Aptamer Conjugation Solve in Real Projects?
Aptamer conjugation is used when a project needs more than target binding alone. In practice, research teams often have a high-value aptamer sequence but still need to convert that recognition event into a usable format: a targeted payload, a fluorescent or biotinylated probe, a nanoparticle guidance element, a liposome surface ligand, or a capture reagent for complex samples. Custom conjugation bridges that gap by linking the aptamer to a functional component while preserving the folding, accessibility, and performance of the binding domain.
This approach is especially valuable when conventional labeling disrupts affinity, when random surface coupling produces inconsistent orientation, or when a carrier system requires controlled ligand density rather than simple adsorption. Aptamer conjugation helps translate sequence-level specificity into practical research outputs such as cell-targeting constructs, nanoparticle functionalization, pull-down reagents, imaging probes, surface-immobilized sensors, and dual-function molecular assemblies designed for defined workflows.
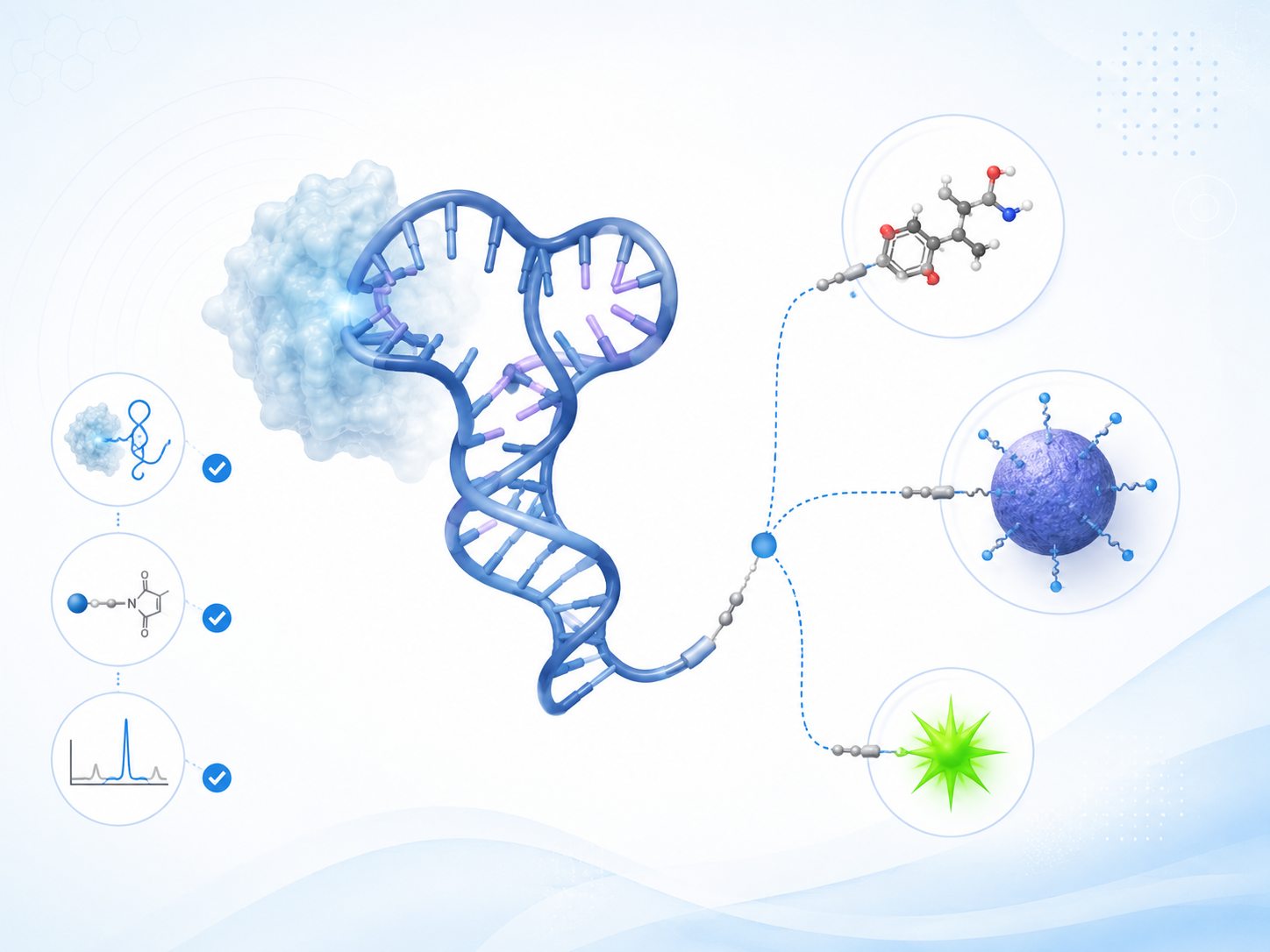 Schematic overview of aptamer conjugation converting sequence-specific binding into payload delivery, surface functionalization, and assay-ready molecular tools.
Schematic overview of aptamer conjugation converting sequence-specific binding into payload delivery, surface functionalization, and assay-ready molecular tools.Where Aptamer Conjugation Programs Commonly Stall
A promising aptamer can lose affinity when a fluorophore, drug, polymer, or carrier is attached too close to a folding-critical region. We help evaluate conjugation site, spacer length, and steric burden so the recognition domain remains accessible after modification.
Random coupling may generate heterogeneous display, reduced target access, or variable batch performance. We design handle placement and coupling routes that improve orientation control, surface presentation, and reproducibility for aptamer-functionalized materials.
Aptamer conjugates can fail because of nuclease sensitivity, linker instability, aggregation, or loss of structure during storage and use. Our workflows consider sequence modifications, buffer/refolding conditions, spacer selection, and project-specific stability checks.
Many teams can make a conjugate but struggle to prove what fraction is correctly assembled, how much free aptamer or free payload remains, and whether binding is retained. We align purification and analytical readouts with the intended use so the final material is not just synthesized, but understood.
Our Aptamer Conjugation Services
We provide custom aptamer conjugation services for research and development programs requiring defined chemistry, preserved aptamer function, and application-relevant analytical support. Beyond standalone aptamer projects, we also support adjacent oligonucleotide bioconjugation needs and project-specific routes informed by bioorthogonal click chemistry when orthogonality and mild reaction conditions are important.
 Aptamer–Drug and Small-Molecule Conjugation
Aptamer–Drug and Small-Molecule Conjugation
Capabilities include:
- Site-conscious attachment at 5′, 3′, or designated internal handles based on aptamer folding and use case
- Support for amine-, thiol-, azide-, alkyne-, or other preinstalled reactive groups
- Linker selection for stable, cleavable, spacer-enabled, or hydrophobicity-managed constructs
- Conjugation of aptamers to small molecules, cytotoxic surrogates, imaging agents, or screening payloads
- Purification strategies selected for free payload removal and product enrichment
- Identity and conjugation confirmation using project-appropriate chromatographic and spectrometric methods
Typical applications:
Targeted uptake studies, payload screening, competitive binding assays, and aptamer-enabled delivery construct development
 Aptamer-Conjugated Nanoparticles and Beads
Aptamer-Conjugated Nanoparticles and Beads
Capabilities include:
- Surface functionalization of aptamers onto metallic, magnetic, polymeric, or silica-based platforms
- Coupling route design for carboxyl, amine, thiol, maleimide, azide, alkyne, or gold-binding surfaces
- Spacer and ligand-density optimization to improve accessibility and reduce steric crowding
- Control of aptamer display for targeting, capture, separation, or sensing workflows
- Support for colloidal stability review and particle-compatible purification logic
- Characterization planning for size distribution, conjugation efficiency, and functional performance
Typical applications:
Targeted nanoplatforms, magnetic capture systems, aptamer-guided biosensors, and functional screening materials
 Aptamer Liposome and Vesicle Surface Conjugation
Aptamer Liposome and Vesicle Surface Conjugation
Capabilities include:
- Aptamer installation through lipid anchors, covalent coupling, or post-insertion style strategies
- Selection of spacer architecture to balance surface exposure with membrane compatibility
- Design support for mono- or multi-ligand presentation on vesicle systems
- Evaluation of density-related effects on binding, aggregation, and accessibility
- Integration of aptamer surface design with liposome composition and downstream workflow needs
- Analytical planning for particle size, conjugation status, and formulation compatibility
Focus areas:
Targeted vesicle engineering, ligand-directed delivery models, and surface-functionalized liposomal research systems
 Aptamer Labeling, Reporter, and Hybrid Conjugates
Aptamer Labeling, Reporter, and Hybrid Conjugates
Capabilities include:
- Fluorophore, quencher, and affinity-tag installation aligned with fluorescence labeling of oligonucleotides and biotin-labeled oligonucleotides workflows
- Aptamer conjugation to complementary oligos, proteins, peptides, or modular reporter handles
- Dual-function construct design for recognition plus signal generation or capture
- Sequence-aware labeling to reduce quenching artifacts or folding disruption
- Purification plans for reporter-labeled conjugates with minimized unconjugated fractions
- Application-focused review for pull-down, imaging, surface assays, and proximity detection
Deliverables:
Aptamer conjugates suitable for assay development, biosensing, enrichment, imaging, and customized molecular tool generation
Key Design Parameters for Aptamer Conjugation
Successful aptamer conjugation depends on more than choosing a reaction handle. Projects usually perform best when aptamer structure, conjugation site, linker behavior, carrier format, and analytical goals are considered together from the beginning. The table below summarizes the design variables most often driving performance and reproducibility.
| Design Parameter | Common Options | Why It Matters | Typical Decision Criteria | Service Relevance |
| Aptamer Format | DNA aptamer, RNA aptamer, truncated aptamer, chemically modified aptamer | Sequence composition and structure affect folding, stability, and conjugation compatibility | Target class, intended medium, storage needs, and downstream assay or carrier format | Determines synthesis route, refolding logic, and modification strategy |
| Conjugation Site | 5′ end, 3′ end, or internal functional handle | Site placement can influence aptamer folding, target access, and steric exposure | Known structure motifs, binding region sensitivity, and required orientation | Critical for preserving affinity after conjugation |
| Reactive Handle | Amine, thiol, azide, alkyne, maleimide partner, biotin, disulfide-forming group | Handle selection defines which chemistry can be used and under what conditions | Payload chemistry, orthogonality needs, and tolerance to aqueous reaction conditions | Drives coupling efficiency and downstream purification difficulty |
| Linker Behavior | Short spacer, PEG spacer, stable linker, reducible linker, acid- or trigger-sensitive linker | Linker design influences flexibility, release profile, hydrophilicity, and steric separation | Whether the conjugate should remain intact, release cargo, or display the aptamer at a distance | Major determinant of function in drug, nanoparticle, and surface applications |
| Carrier or Payload Class | Small molecule, fluorophore, protein/peptide, nanoparticle, bead, liposome, complementary oligo | Different payloads introduce different steric, solubility, and analytical challenges | Intended use, loading requirement, and compatibility with aptamer structure | Shapes the reaction route and functional assay plan |
| Valency or Surface Density | Single aptamer, multivalent display, low-density surface loading, high-density surface loading | Excess density can impair accessibility or promote aggregation; too little can reduce performance | Nanoparticle size, surface area, target abundance, and required avidity | Important for reproducible particle and surface-functionalization outcomes |
| Purification & QC Plan | HPLC/UPLC, desalting, SEC, ultrafiltration, gel analysis, particle cleanup workflows | Product quality depends on removal of free aptamer, free payload, and side products | Molecular size, formulation, and final use requirements | Ensures the final conjugate is suitable for experimental use rather than nominally reacted only |
Aptamer Conjugation Chemistries and When to Use Them
There is no single best chemistry for every aptamer conjugation program. The right route depends on the reactive groups already present, the sensitivity of the aptamer sequence, the nature of the payload or carrier, and whether the final construct must be permanent, reversible, or highly orthogonal to other functionalities.
| Conjugation Strategy | Typical Handle Pair | Best Fit | Key Considerations |
| Amide Coupling | Amine + activated carboxyl group | Small molecules, polymers, proteins, and surface functionalization workflows | Broadly accessible but requires control of side reactions and regioselectivity |
| Thiol–Maleimide Coupling | Thiol + maleimide | Aptamer–payload conjugates, protein interfaces, and nanoparticle attachment | Useful for directional conjugation when thiol placement is well defined |
| Azide–Alkyne Click Chemistry | Azide + alkyne or cyclooctyne partner | Orthogonal assembly of aptamer conjugates in multifunctional systems | Well suited when clean coupling and minimal interference with other groups are required |
| Disulfide or Redox-Sensitive Coupling | Thiol-containing partners or reducible linkers | Constructs requiring controlled cleavage or reversible linkage logic | Link stability must match storage and experimental environment |
| Biotin–Affinity Assembly | Biotin + streptavidin/avidin platform | Capture assays, pull-down systems, immobilization, and rapid modular assembly | Strong interaction and easy formatting, but complex size and valency may affect presentation |
| Surface-Specific Anchoring | Gold–thiol, silica/polymer surface handles, bead-coupling groups | Aptamer-functionalized nanoparticles, magnetic beads, electrodes, and biosensor surfaces | Surface density, orientation, and passivation must be balanced for functional display |
Analytical Characterization and Quality Control for Aptamer Conjugates
Aptamer conjugation projects benefit from analytical planning that matches the final format. A labeled aptamer probe, a drug conjugate, and an aptamer-functionalized nanoparticle do not require the same verification package. We structure characterization around identity, conjugation status, residual unconjugated species, and retained function in the intended workflow.
| Analytical Category | Typical Methodology | Purpose in the Project | Data Delivered |
| Sequence / Identity Confirmation | LC-MS, MALDI-MS, or other mass-confirmation workflows as appropriate | Verifies the expected aptamer and confirms successful installation of the designed modification or payload | Molecular weight confirmation and identity summary |
| Conjugation Efficiency & Purity | HPLC, UPLC, gel-based analysis, SEC, or formulation-compatible cleanup assessment | Distinguishes target conjugate from free aptamer, free payload, and partially reacted species | Purity profile, chromatograms, and product-distribution summary |
| Residual Free Component Assessment | Orthogonal chromatographic or spectroscopic readouts selected by conjugate class | Confirms that unbound label, cargo, or carrier-associated contaminants are adequately reduced | Residual component analysis and cleanup evaluation |
| Binding Retention Check | Project-relevant binding assays such as plate-based formats, SPR/BLI-style evaluation, or cell-binding analysis | Determines whether the aptamer still performs after conjugation and purification | Comparative binding or functional performance data |
| Particle / Surface Characterization | Size-distribution, surface-property, or morphology methods where relevant to nanoparticle and liposome programs | Evaluates whether conjugation changed particle behavior, aggregation tendency, or display characteristics | Particle characterization summary for formatted systems |
| Stability-Oriented Review | Storage-condition checks, buffer evaluation, and follow-up analysis under defined use conditions | Identifies degradation, loss of signal, linker instability, or structure-sensitive failure modes | Stability observations and recommended handling conditions |
| Batch Comparability | Comparative purity, identity, and functional review across prepared lots | Helps maintain reproducibility for ongoing assay, platform, or material-development work | Lot comparison summary and consistency review |
Workflow for Custom Aptamer Conjugation Projects
We begin by reviewing the aptamer format, known binding behavior, intended target, and final use scenario. This stage identifies folding-sensitive regions, preferred termini for modification, and any constraints introduced by the payload, surface, or formulation.
Based on the project goal, we select a chemistry strategy and spacer architecture that balances coupling efficiency with aptamer accessibility. Stable, cleavable, orthogonal, or surface-specific routes can be matched to the intended system.
We prepare the aptamer and conjugation partner with the required reactive groups or formatted surfaces. This may include terminal modifications, partner activation, carrier surface conditioning, or ligand-density planning before the coupling step.
The conjugation is performed under conditions selected for the aptamer and conjugation partner, followed by purification designed to remove unconjugated species and reaction byproducts. The purification route is chosen according to molecular size and final format.
We confirm the assembled conjugate using application-relevant analytical methods and, where needed, evaluate retained binding or performance in an appropriate assay format. This helps ensure the material is suitable for real downstream use rather than only structurally plausible.
If the first design needs refinement, we can iterate on conjugation site, linker, density, or purification logic. Final deliverables typically include the conjugate, analytical data, and practical recommendations for handling, storage, and downstream integration.
Why Teams Choose Our Aptamer Conjugation Platform
We do not treat aptamers as generic oligos. Conjugation routes are selected with attention to folding, terminus sensitivity, target accessibility, and the practical consequences of payload size or surface presentation.

Our support extends across direct payload conjugates, aptamer-labeled probes, nanoparticle and bead functionalization, liposome surface engineering, and hybrid constructs that combine recognition with signal or cargo functions.
We align QC with how the conjugate will actually be used. That means focusing not only on identity and purity, but also on free-component removal, retained binding, density-related effects, and formatted-system behavior when required.
Aptamer projects often intersect with peptide, protein, nanoparticle, or reporter chemistry. Our workflows are built to integrate those interfaces efficiently, including related needs such as protein–oligonucleotide conjugation and modular assay probe design.
Research Applications of Aptamer Conjugation
Aptamer-Directed Payload Delivery Models
- Small-molecule, oligo, or reporter payload attachment for targeted uptake and functional studies.
- Linker-enabled constructs for comparing stable versus releasable architectures.
- Screening systems that connect molecular recognition with defined cargo presentation.
Nanoparticle, Bead, and Liposome Functionalization
- Surface display of aptamers for targeting, enrichment, or multivalent binding studies.
- Magnetic and optical materials formatted for capture, separation, or sensor development.
- Vesicle and nanocarrier systems requiring controlled ligand density and orientation.
Imaging, Detection, and Reporter Probes
- Fluorescent, quenched, or biotinylated aptamer probes for assay and imaging workflows.
- Dual-function constructs that combine target recognition with signal generation.
- Customized probes for plate-based assays, microscopy-adjacent workflows, and biosensing.
Capture, Immobilization, and Surface-Based Assays
- Aptamer-functionalized supports for pull-down, sorting, and affinity capture.
- Surface immobilization strategies for electrodes, chips, and biosensor interfaces.
- Structured formats for target detection, interaction analysis, and analytical tool development.
Move Your Aptamer Conjugation Project Forward with a Defined Chemistry and QC Strategy
Whether you are developing an aptamer–drug construct, functionalizing a nanoparticle surface, preparing a labeled probe, or building a custom capture reagent, we provide project-specific support from conjugation design to purified material and analytical review.
Our team works with research and development groups to reduce avoidable trial-and-error in aptamer formatting, handle installation, linker selection, and conjugate verification so that each construct is matched to its downstream application rather than treated as a generic modification exercise.Contact our scientific team to discuss your aptamer conjugation goals and explore a workflow tailored to your sequence, payload, carrier, and analytical requirements.
Frequently Asked Questions (FAQ)
Aptamers can be conjugated to small molecules, fluorophores, quenchers, biotin, peptides, proteins, complementary oligos, nanoparticles, beads, liposomes, and other surface-formatted materials depending on the project goal.
The best site is usually the terminus or internal handle that minimizes disruption to folding and target access. Sequence structure, known binding motifs, and the size of the conjugated partner all matter.
Common options include amide coupling, thiol–maleimide coupling, azide–alkyne click chemistry, biotin-based affinity assembly, disulfide-style linkages, and surface-specific anchoring such as gold–thiol coupling.
Binding is protected by choosing a suitable modification site, using an appropriate spacer, controlling reaction conditions, and confirming function after purification with a project-relevant binding assay.
Yes. Aptamers are often installed on nanoparticles, beads, and liposomes as targeting or capture ligands, but ligand density, orientation, and colloidal stability need to be controlled carefully.

