Bioorthogonal Click Chemistry in Biochemical Research and Drug Discovery
Bioorthogonal click chemistry has found numerous applications in chemical biology, bioimaging, and drug discovery. By incorporating clickable groups into biomolecules of interest and then reacting them with small molecule probes or imaging agents, selectively tagging and visualization of specific molecules within living systems can be achieved. This technique allows for the study of biological processes with high spatial and temporal resolution.
Metabolic Engineering and Cell Imaging
Metabolic engineering is a strategy that utilizes the inherent enzymatic machinery of cells to introduce non-natural functional groups into target biomolecules. Metabolites, also known as chemical reporters, containing bioorthogonal functional groups, participate in metabolic pathways during biosynthesis and are incorporated into the final target biomolecules. This strategy allows the reporters to be recognized by another component in the bioorthogonal reaction, leading to the introduction of fluorescent labels.
Currently, this approach is used for imaging proteins, polysaccharides, and other intracellular biomolecules (Fig. 1). It provides a new research avenue for detecting biomolecules that is challenging to achieve through genetic manipulation. Typically, azide and alkyne groups are used for metabolite labeling because of their small size and tolerance to various cellular enzymes. CuAAC or SPAAC involving these groups have been employed for metabolite labeling. For example, the SPAAC successfully achieved fluorescent labeling of cell surface polysaccharides for subsequent cell imaging.
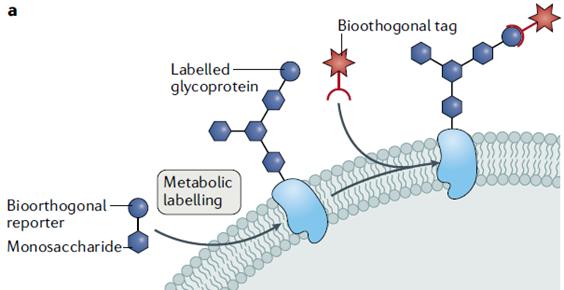 Fig. 1 Metabolite Labeling through Bioorthogonal Coupling (Scinto, 2021)
Fig. 1 Metabolite Labeling through Bioorthogonal Coupling (Scinto, 2021)
Genetic Code Expansion
Genetic code expansion allows the incorporation of non-natural amino acids into proteins, providing reactive sites for bioorthogonal reactions. Genetic code expansion relies on the use of orthogonal aminoacyl-tRNA synthetase (aaRS)-tRNA pairs to enable the co-translational incorporation of non-natural amino acids at the amber stop codon that refers to the UAG codon. Orthogonal means that it does not interfere with the normal function of endogenous aaRS-tRNA pairs.
Over the past 20 years, scientists have developed many orthogonal aaRS-tRNA pairs for incorporating non-natural amino acids with bioorthogonal functionality into bacteria, yeast, and mammalian cells, enabling site-specific modifications and protein labeling.
- The non-natural amino acid is carried by a tRNA generated by orthogonal aaRS. This tRNA recognizes the UAG.
- During translation of the new protein, the original UAG stop codon does not terminate translation but incorporates the non-natural amino acid into the protein, allowing further translation.
- The translation process is terminated by other stop codons, resulting in the incorporation of the non-natural amino acid into the protein.
Commonly used non-natural amino acids often contain functional groups such as ketones, azides, alkynes, and tetrazines, which can undergo bioorthogonal reactions. This method enables protein surface modifications, site-specific conjugation of antibody-drug conjugates, protein fluorescent labeling, and other processes.
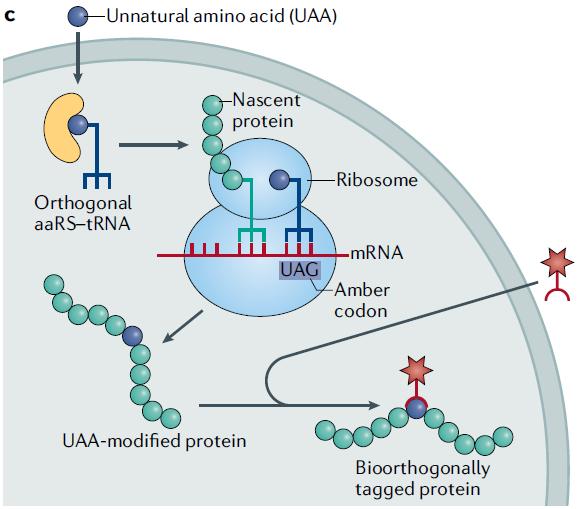 Fig. 2 Principle of Genetic Code Expansion (Scinto, 2021)
Fig. 2 Principle of Genetic Code Expansion (Scinto, 2021)
Drug Discovery
The application of bioorthogonal reactions in the field of drug discovery is becoming increasingly widespread, including target identification (such as phenotype screening), drug-target binding studies, and selective screening of lead compounds.
Application of bioorthogonal reactions in ABPP
In drug discovery, activity-based protein profiling (ABPP) is a valuable method that can distinguish active proteins from inactive proteins (such as those derived from zymogens, post-translational modifications, or inhibitor binding forms) in complex physiological systems. ABPP has various applications, including identifying cell viability and abnormal enzyme activity, providing strong support for drug discovery. ABPP relies on probes with a reactive warhead, linker, and label for detection. When using probes for direct detection, the structure of the probe may affect cell permeability and cellular localization. Therefore, scientists have developed ABPP techniques based on bioorthogonal reactions.
- A small fragment containing an alkyne or azide group is first combined with the target protein, which has a smaller size and minimal impact on the cellular environment.
- Another fluorescent label capable of participating in a click reaction is added to bind with the previous fragment, enabling subsequent detection.
Application of bioorthogonal reactions in ADC
Another application of bioorthogonal reactions is the development of antibody-drug conjugates (ADCs). For example, the CL2A linker in Sacituzumab govitecan contains a triazole structure, indicating the involvement of a click reaction in the preparation of the CL2A linker. In addition, some site-specific conjugation techniques in ADCs also incorporate the previously mentioned genetic code expansion technology. The antibodies containing non-natural amino acids can undergo bioorthogonal reactions with toxins, enabling site-specific conjugation of ADCs.
Bioorthogonal Uncaging
Bioorthogonal uncaging refers to the release of payloads by cleaving small molecular fragments using bioorthogonal reactions. In 2006, the first example of bioorthogonal uncaging using the Staudinger reaction to disrupt the structure of aminoformate esters and release drug payloads has been reported. In additon, other Bioorthogonal reactions, cuch as tetrazine ligation, also can be used for payload uncaging.
Bioorthogonal uncaging can be combined with imaging techniques. For example, the in vivo studies of bioorthogonal uncaging of ADCs. When most of the ADCs reached the surface of tumor cells but had limited internalization, radiolabeled probes with tetrazine structures were introduced, cleaving the linker and releasing the toxin. Meanwhile, the portion containing the radionuclide was connected to the antibody through the orthogonal reaction, allowing the subsequent observation of ADC's biodistribution through imaging technique.
Conclusion
The application of bioorthogonal reactions is not only evident in precise and traceless compound synthesis but also in various fields, including metabolic engineering and imaging, genetic code expansion, drug discovery, and drug release. Furthermore, bioorthogonal reactions have promising applications in polymer chemistry, materials science, 3D printing, and other areas. It is expected that bioorthogonal reactions will continue to play an important role in different disciplines and interdisciplinary fields in the future.
Frequently Asked Questions (FAQ)
Bioorthogonal click chemistry refers to chemical reactions that occur selectively between functional groups in living systems without interfering with natural biological processes. It allows researchers to tag, track, and study biomolecules with high precision, enabling advancements in various scientific fields, including chemical biology, bioimaging, and drug discovery.
In metabolic engineering, bioorthogonal click chemistry is used to introduce non-natural functional groups into biomolecules through metabolic pathways. These functional groups can be recognized by bioorthogonal reactions, allowing the labeling of proteins, lipids, or other intracellular biomolecules for detailed imaging and tracking within living cells.
Genetic code expansion enables the incorporation of non-natural amino acids into proteins by using orthogonal tRNA-synthetase pairs. These non-natural amino acids carry bioorthogonal functional groups that can undergo specific reactions, providing new tools for protein labeling and targeted modifications, enabling more precise control over protein function.
ABPP is a method used in drug discovery to identify active proteins in a system by tagging them with bioorthogonal probes. These probes contain functional groups like azides or alkynes, which react with bioorthogonal partners for detection. This allows researchers to differentiate between active and inactive proteins in living cells without affecting their natural function.
Yes, bioorthogonal click chemistry is widely used in cellular imaging to label specific biomolecules like proteins, lipids, or polysaccharides. By incorporating bioorthogonal groups into these molecules, researchers can visualize their behavior in real-time within live cells, offering insights into cellular dynamics and biomolecular interactions.


Reference
- Scinto, S. L., et al., Bioorthogonal chemistry, Nat. Rev. Methods Primers, 2021, 1, 29.