Fluorescently Labeled Nucleotide
Custom Nucleotide ConjugationBase-modified NucleotidesValidated Reagents for NGS, qPCR & Molecular Diagnostics
Enable robust fluorescence-based detection with custom fluorescently labelled nucleotides manufactured for enterprise R&D, assay development, and regulated workflows. We supply labelled dNTPs and nucleotide analogs (e.g., fluorescent dUTP/dCTP variants), as well as application-matched formulations for enzymatic incorporation in PCR, reverse transcription, DNA polymerase extension, and library preparation. Our production integrates controlled conjugation chemistry, purification designed for nucleotide reagents, and quality verification aligned with the needs of biotechnology, pharmaceutical, diagnostics, and CRO/CDMO partners.
Whether you are optimizing qPCR probe systems, developing fluorescence-based sequencing workflows, building diagnostic assays, or running high-throughput screening platforms that rely on polymerase-compatible fluorophores, we support project-specific design: fluorophore selection by excitation/emission requirements, incorporation compatibility (polymerase and buffer conditions), and impurity control to reduce background and inhibition risk. Deliverables include the labelled nucleotide product and documentation to support method development and tech transfer.
What Are Fluorescently Labelled Nucleotides?
Fluorescently labelled nucleotides are nucleoside triphosphates (or nucleotide analogs) functionalized with a fluorophore so they can be detected by fluorescence during or after enzymatic incorporation into DNA or RNA. They are widely used in PCR and qPCR workflows, nucleic-acid labeling and hybridization assays, imaging of nucleic-acid processes in vitro, and sequencing and library-prep methods that require fluorescent readouts. By tailoring the label position, linker chemistry, and purification strategy to the nucleotide and the intended enzyme system, labelled nucleotides can be used to generate measurable signal while maintaining practical incorporation efficiency for assay development and research applications.
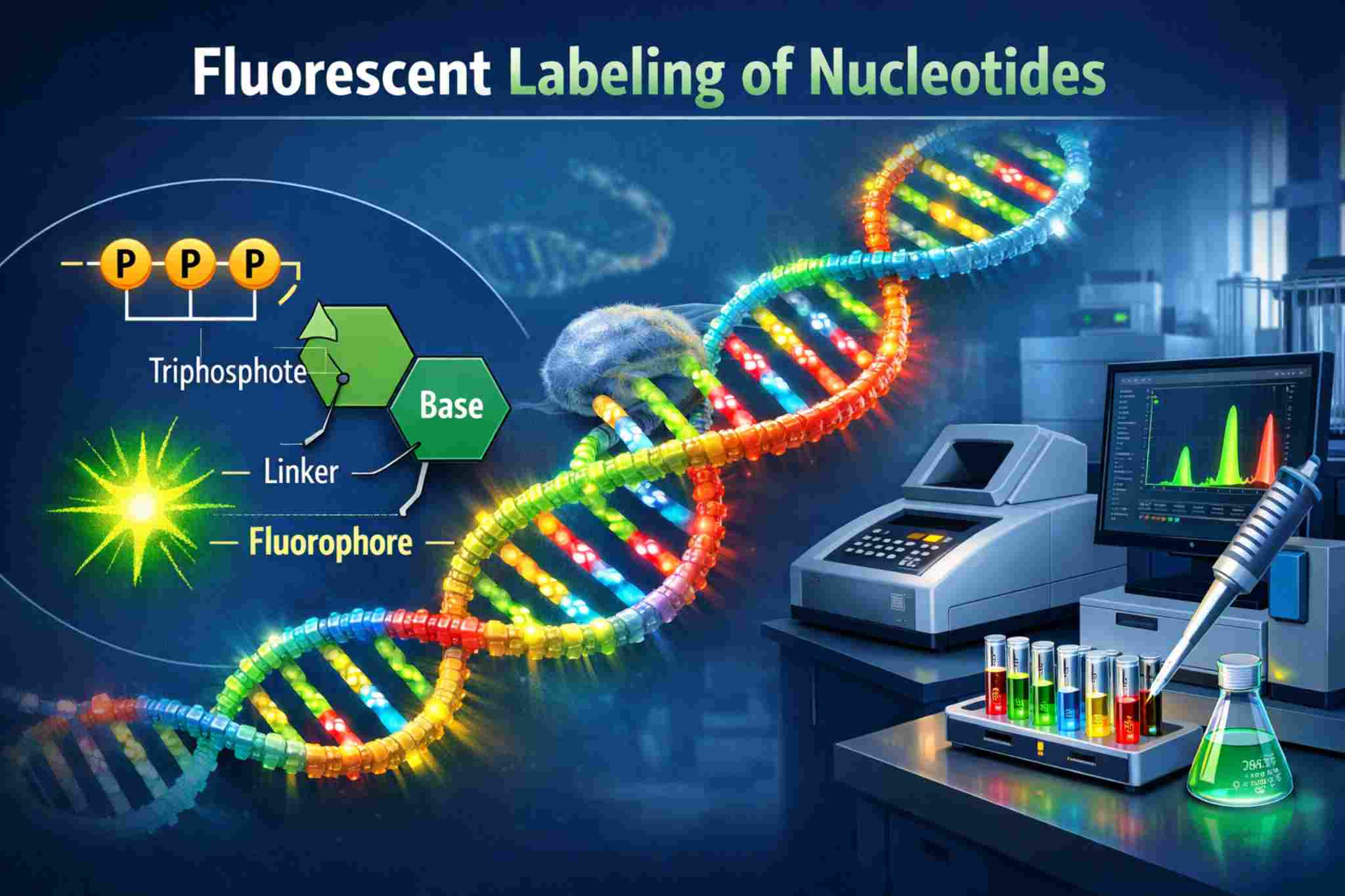 Illustration of fluorescently labeled nucleotide structure, fluorophore linkage, and incorporation into DNA for multiplex qPCR and sequencing applications.
Illustration of fluorescently labeled nucleotide structure, fluorophore linkage, and incorporation into DNA for multiplex qPCR and sequencing applications.Enterprise Challenges in Fluorescently Labelled Nucleotide Development
Bulky fluorophores or poorly positioned linkers can reduce DNA polymerase or reverse transcriptase compatibility. We evaluate fluorophore size, linker length, and labeling position to improve practical incorporation performance in PCR, RT-PCR, and extension assays used in diagnostics and sequencing workflows.
Residual free dye, partially hydrolyzed nucleotides, or side products can elevate background signal and compromise assay sensitivity. Our purification strategies focus on removing unreacted fluorophores and closely related impurities to support low-noise qPCR, hybridization, and imaging assays.
Inconsistent dye-to-nucleotide ratios and purity profiles disrupt assay validation and tech transfer. We implement controlled synthesis parameters and standardized analytical verification to support reproducible performance for enterprise customers scaling from R&D to pilot production.
Impurities, metal contamination, or suboptimal buffer systems may inhibit polymerases, affecting amplification efficiency and sequencing read quality. Our manufacturing and QC workflows are aligned with molecular biology applications to minimize inhibitory components.
Our Enterprise Fluorescently Labelled Nucleotide Services
We provide custom and project-based manufacturing of fluorescently labelled nucleotides tailored to molecular diagnostics developers, NGS technology providers, biotechnology companies, and CRO/CDMO partners. Our services focus on application-matched design, analytical verification, and scalable production workflows to support assay development, platform validation, and commercial preparation.
 Fluorescent dNTP & Nucleotide Analog Synthesis
Fluorescent dNTP & Nucleotide Analog Synthesis
Capabilities include:
- Synthesis of fluorescently labelled dUTP, dCTP, and other nucleotide analogs for enzymatic incorporation
- Selection of fluorophores compatible with common qPCR and sequencing detection channels (e.g., green, orange, red, near-IR ranges)
- Linker optimization to balance fluorescence intensity and polymerase compatibility
- Adaptation for PCR, RT-PCR, nick translation, and primer extension applications
- Support for research-use and assay-development grade material
- Purification using chromatographic methods suitable for nucleotide reagents
- Analytical characterization including HPLC profiling and spectral verification
- Small- to mid-scale production for platform evaluation and pilot studies
- Technical consultation for integration into molecular diagnostic workflows
- Documentation package to facilitate internal validation and method development
Common fluorophore families:
Fluorescein derivatives (FAM-type), HEX-type dyes, ROX-type dyes, Cy3/Cy5 families, selected near-IR dyes compatible with real-time PCR and fluorescence detection systems
 Multiplex Assay & Platform Integration Support
Multiplex Assay & Platform Integration Support
Capabilities include:
- Multi-color nucleotide panel design for multiplex qPCR and fluorescence-based assays
- Spectral compatibility assessment aligned with major qPCR instrument filter sets
- Guidance on minimizing spectral overlap and signal cross-talk
- Compatibility evaluation with commonly used DNA polymerases and reverse transcriptases
- Optimization strategies for incorporation ratios and signal balance
- Support for NGS library preparation workflows requiring fluorescent detection
- Risk assessment for assay sensitivity and limit-of-detection considerations
- Batch reproducibility planning for tech transfer to manufacturing partners
- Stability testing under defined storage and shipping conditions
- Collaborative project management for enterprise R&D and diagnostic development teams
Application focus:
Molecular diagnostics, infectious disease testing, oncology biomarker assays, NGS platform development, and high-throughput screening systems
 qPCR & Molecular Diagnostics Nucleotide Support
qPCR & Molecular Diagnostics Nucleotide Support
Capabilities include:
- Development of fluorescently labelled dUTP and related nucleotide analogs for enzymatic incorporation in PCR and RT-PCR workflows
- Fluorophore selection aligned with common real-time PCR instrument detection channels (e.g., FAM-, HEX-, ROX-, Cy5-compatible ranges)
- Linker optimization to balance fluorescence intensity with practical polymerase incorporation efficiency
- Impurity profiling and purification strategies to reduce polymerase inhibition and background fluorescence
- Multiplex panel support to minimize spectral overlap in multi-target diagnostic assays
- Stability evaluation under defined storage and transport conditions relevant to kit manufacturing
- Batch reproducibility planning to support assay validation and technology transfer
- Documentation package including chromatographic purity data and spectral characterization
- Support for RUO assay development and pre-commercial diagnostic workflow optimization
- Collaborative technical consultation with assay development teams
Typical applications:
Infectious disease testing, oncology biomarker detection, multiplex pathogen panels, and fluorescence-based amplification assays
 NGS & Fluorescence-Based Platform Development Support
NGS & Fluorescence-Based Platform Development Support
Capabilities include:
- Custom fluorescent nucleotide analog development for sequencing-related enzymatic workflows
- Evaluation of compatibility with engineered DNA polymerases and reverse transcriptases
- Spectral optimization for multi-channel fluorescence detection systems
- Linker engineering to reduce steric hindrance and maintain enzyme turnover efficiency
- Controlled purification to remove free dye and hydrolyzed triphosphates that may impact read fidelity
- Support for fluorescence-based extension assays and library preparation development
- Analytical characterization including HPLC purity profiling and UV-Vis spectral confirmation
- Feasibility-scale to pilot-scale production for platform validation programs
- Long-term supply planning for platform developers transitioning from R&D to commercial preparation
- Cross-functional project coordination with enzyme engineering and assay integration teams
Application focus:
Next-generation sequencing workflow development, fluorescence-based enzymatic assays, synthetic biology systems, and high-throughput nucleic acid analysis platforms
Fluorophores Optimized for Fluorescently Labelled Nucleotides
Selection of fluorophores for nucleotide conjugation requires careful consideration of polymerase compatibility, spectral performance, photostability, and instrument filter alignment. The following dyes are widely used in fluorescent dNTPs, qPCR incorporation assays, and fluorescence-based sequencing and detection workflows.
| Fluorophore Family | Typical Excitation (nm) | Typical Emission (nm) | Common Nucleotide Applications | Technical Considerations |
| FAM-Type (Fluorescein Derivatives) | 495 | 520 | qPCR incorporation assays, fluorescent dUTP labeling, amplification tracking | High brightness, widely compatible with real-time PCR systems; moderate photostability |
| HEX / VIC-Type Dyes | 535 | 555 | Multiplex qPCR assays, dual-channel amplification systems | Improved spectral separation from FAM; suitable for multiplex diagnostics |
| TAMRA / ROX-Type Dyes | 560–585 | 580–610 | Reference normalization channels, multiplex amplification assays | Good photostability; commonly aligned with ROX reference channels in qPCR instruments |
| Cy3 Family | 550 | 570 | Fluorescent nucleotide incorporation, hybridization detection assays | Bright emission; suitable for imaging and fluorescence-based enzymatic assays |
| Cy5 Family | 640–650 | 660–670 | Multiplex PCR, NGS-associated fluorescence workflows | Low background autofluorescence; ideal for red/near-IR detection channels |
| Near-Infrared (NIR) Dyes | 750–780 | 770–810 | Advanced multiplex systems, specialized detection platforms | Reduced biological autofluorescence; useful for high-sensitivity detection systems |
Nucleotide Conjugation Chemistry & Engineering Strategies
Fluorescent nucleotide design requires controlled conjugation strategies that preserve triphosphate stability and polymerase compatibility. Our platform integrates nucleotide-specific attachment chemistries and linker engineering approaches to balance fluorescence performance with enzymatic incorporation efficiency.
| Conjugation Strategy | Chemistry Basis | Common Applications | Technical Advantages |
| Base-Modified Nucleotide Labeling | Fluorophore attachment to modified nucleobases (commonly uracil or cytosine analogs) | Fluorescent dUTP incorporation, PCR tracking, labeling assays | Maintains triphosphate integrity while allowing enzyme-compatible labeling |
| Linker-Assisted Dye Conjugation | Flexible carbon or PEG-based spacers between dye and nucleotide base | qPCR multiplex assays, fluorescence imaging workflows | Reduces steric hindrance and improves polymerase incorporation efficiency |
| Click-Chemistry–Enabled Nucleotide Modification | Azide-alkyne cycloaddition for post-synthetic dye attachment | Custom nucleotide analog development, research-use platforms | High specificity; enables modular dye exchange strategies |
| Enzyme-Compatible Triphosphate Stabilization | Controlled reaction and purification conditions to preserve triphosphate structure | PCR, RT-PCR, extension assays, sequencing-related workflows | Minimizes hydrolysis and protects polymerase activity |
| Spectral Panel Engineering | Fluorophore selection based on excitation/emission separation | Multiplex diagnostic assays, multi-channel detection platforms | Reduces cross-talk and improves signal discrimination |
Quality Control & Analytical Validation for Fluorescent Nucleotides
Fluorescently labelled nucleotides require rigorous analytical validation to ensure polymerase compatibility, spectral reliability, and batch reproducibility. Our QC framework is aligned with molecular diagnostics and sequencing workflow requirements.
| QC Parameter | Analytical Method | Delivered Data |
| Chemical Purity | HPLC (ion-exchange or reverse-phase) | Chromatogram, % purity profile |
| Molecular Identity Confirmation | LC-MS or high-resolution MS | Mass confirmation report |
| Triphosphate Integrity | HPLC stability assessment | Hydrolysis monitoring data |
| Fluorescence Spectral Characterization | UV-Vis and fluorescence spectroscopy | Excitation/emission spectra, absorbance ratios |
| Free Dye Residual Analysis | Chromatographic separation | Free dye percentage quantification |
| Polymerase Compatibility Screening (Optional) | Incorporation assay with representative polymerases | Relative incorporation performance data |
| Stability Testing | Controlled temperature storage study | Degradation and fluorescence retention profile |
Development Workflow for Fluorescently Labelled Nucleotides
We evaluate the intended workflow (qPCR, RT-PCR, NGS-related enzymatic assays, or fluorescence-based detection systems), target polymerase, detection instrument channels, and multiplexing requirements. This ensures fluorophore selection and nucleotide modification strategy are aligned with real assay conditions.
Fluorophore attachment strategy (commonly base-modified analogs such as dUTP derivatives) and linker design are optimized to balance fluorescence intensity with enzymatic incorporation efficiency, minimizing steric hindrance that may impact polymerase activity.
Conjugation conditions are carefully managed to preserve triphosphate integrity and prevent hydrolysis. Reaction parameters are selected to maintain chemical stability while achieving defined dye-to-nucleotide composition.
Ion-exchange or reverse-phase chromatographic methods are applied to remove free dye, partially hydrolyzed nucleotides, and closely related impurities that could inhibit polymerases or increase assay background.
Each batch undergoes HPLC purity profiling, LC-MS identity confirmation, and fluorescence spectral analysis. Optional polymerase incorporation screening can be conducted under representative assay conditions.
Products are supplied under light-protective, stability-conscious conditions with supporting analytical documentation. Our team provides integration guidance to facilitate assay validation, platform optimization, and reproducibility across batches.
Enterprise Advantages in Fluorescently Labelled Nucleotide Development
Fluorescent nucleotide performance depends heavily on enzyme tolerance. Our development process emphasizes linker engineering and base-modification strategies designed to preserve practical incorporation efficiency in PCR, RT-PCR, and fluorescence-based sequencing workflows.

We align fluorophore selection with common diagnostic instrument filter sets and multi-channel detection systems, supporting multiplex assay development while minimizing spectral overlap and signal cross-talk in enterprise platforms.
Fluorescent nucleotide stability directly impacts assay reliability. Our workflows focus on chromatographic purification, free dye removal, and hydrolysis monitoring to reduce enzyme inhibition risk and ensure consistent analytical performance.
We support enterprise customers from feasibility evaluation to pilot-scale production, enabling smooth transition from R&D optimization to larger-batch supply while maintaining defined analytical specifications.
Applications in Molecular Diagnostics & Sequencing Technologies
Real-Time PCR & Multiplex Diagnostics
- Incorporation-based fluorescence tracking in qPCR amplification workflows.
- Multi-channel multiplex pathogen and biomarker detection panels.
- Signal normalization strategies aligned with reference dye systems.
- Support for assay sensitivity optimization and background reduction.
Next-Generation Sequencing Workflow Development
- Fluorescent nucleotide analogs for enzymatic extension and detection workflows.
- Spectral engineering for multi-channel fluorescence detection systems.
- Compatibility evaluation with engineered polymerases used in sequencing platforms.
- Support for library preparation and fluorescence-based quality control assays.
Oncology & Infectious Disease Testing
- Detection of low-abundance mutations and pathogen-specific nucleic acids.
- Multiplex fluorescence systems for simultaneous target amplification.
- Integration into fluorescence-based amplification and hybridization assays.
- Stability-aligned design for diagnostic kit manufacturing.
Synthetic Biology & Enzyme Engineering
- Evaluation of engineered polymerase tolerance to modified nucleotides.
- Fluorescence-based monitoring of in vitro transcription and DNA synthesis systems.
- Screening platforms for nucleotide analog incorporation efficiency.
- Development of fluorescence-enabled enzymatic assays.
High-Throughput Nucleic Acid Analysis Platforms
- Fluorescent incorporation systems for automated amplification platforms.
- Multi-color detection in large-scale screening systems.
- Signal optimization for low-background fluorescence detection.
- Adaptation to instrument-specific excitation and emission configurations.
What Our Research Clients Say

"We needed fluorescently labeled nucleotides that would integrate cleanly into our multiplex qPCR workflow without elevating background or impacting amplification performance. The batch documentation and analytical data made internal qualification straightforward, and the reagent behavior was consistent across runs during assay development."
— Director of Assay Development, Molecular Diagnostics Company
"Our team required nucleotide analog support aligned to instrument channel requirements and polymerase compatibility constraints. The project communication was technical and efficient, and the delivered material was accompanied by clear purity profiling and spectral verification data that supported rapid iteration in our platform optimization program."
— Principal Scientist, Sequencing Technology Developer
"For kit integration, stability and batch reproducibility were our primary concerns. The supply planning approach and consistent analytical profile reduced re-validation workload and helped our team maintain predictable performance in fluorescence-based amplification assays across multiple internal sites."
— Technical Lead, Reagent & Kit Manufacturer
Partner with Us for Enterprise-Scale Fluorescent Nucleotide Development
Whether you are developing multiplex diagnostic assays, optimizing qPCR workflows, engineering sequencing platforms, or scaling fluorescence-based enzymatic systems, we provide application-driven support tailored to your technical and operational requirements. Our team collaborates with assay developers, platform engineers, and manufacturing groups to align fluorophore selection, nucleotide modification strategy, analytical validation, and scalable supply planning.
Engage with our technical specialists to discuss polymerase compatibility considerations, multiplex spectral design, impurity control strategies, and long-term supply alignment for your program. Contact our scientific team to initiate a confidential project discussion or request a technical consultation.
Frequently Asked Questions (FAQ)
Fluorescently labelled nucleotides are used in applications where fluorescence detection is required during or after enzymatic incorporation into DNA or RNA. Common uses include real-time PCR (qPCR), multiplex diagnostic assays, nucleic acid labeling workflows, fluorescence-based extension assays, sequencing-related detection systems, and enzyme engineering studies. In enterprise settings, they are often integrated into assay platforms rather than used as standalone reagents.
Fluorescent dUTP derivatives are the most commonly used labelled nucleotides for PCR-based applications. Uracil-based analogs are often selected because modification at this position is generally more compatible with DNA polymerase activity compared to bulkier modifications on other bases. The choice of fluorophore (e.g., FAM-, HEX-, ROX-, or Cy5-type dyes) depends on the instrument detection channels and multiplex design.
Fluorescent nucleotides are incorporated directly into newly synthesized DNA or RNA strands during enzymatic reactions. Fluorescent probes (such as hydrolysis probes or hybridization probes) bind to a specific sequence and generate signal through cleavage or conformational change. Fluorescent nucleotide incorporation provides distributed signal throughout amplified products, while probes provide sequence-specific detection.
Certain fluorescent nucleotide analogs are used in fluorescence-based sequencing and enzymatic detection workflows. Compatibility depends on:
Polymerase engineering
Dye position and linker design
Platform-specific detection chemistry
Sequencing applications typically require customized nucleotide engineering rather than standard PCR-grade labelled nucleotides.
Yes, when spectral separation is carefully designed. Multiplex compatibility depends on:
Fluorophore excitation/emission separation
Instrument filter configuration
Minimizing spectral cross-talk
Proper normalization strategy
Enterprise multiplex panels typically use combinations such as FAM + HEX + ROX + Cy5, depending on platform capability.

