Aptamer Gold Nanoparticle Conjugation Services
Precise AuNP Surface FunctionalizationTarget-Responsive Aptamer EngineeringCustom Conjugates for Sensing, Imaging & Delivery Research
Build assay-ready and study-ready aptamer–gold nanoparticle conjugates with a workflow designed for research teams working in biosensing, molecular recognition, cell-targeting studies, and nanomaterial-enabled delivery. Aptamer gold nanoparticle conjugation combines the sequence programmability and target selectivity of aptamers with the plasmonic, surface-addressable, and multivalent properties of gold nanoparticles, making it a practical platform for colorimetric assays, lateral flow probes, SERS/electrochemical systems, and proof-of-concept cargo presentation.
We support custom development from aptamer review and terminal modification strategy through gold nanoparticle selection, conjugation, passivation, purification, and analytical characterization. Projects can be aligned with new build programs or with optimization of existing aptamer-conjugated nanoparticle constructs, and can be integrated with broader oligonucleotide bioconjugation or nucleic acid-functionalized gold nanoparticle probe development needs.
What Problems Can Aptamer Gold Nanoparticle Conjugation Solve?
Many projects fail not because the aptamer sequence is poor, but because the aptamer is attached to gold nanoparticles in a way that compromises folding, accessibility, colloidal stability, or batch reproducibility. Aptamer gold nanoparticle conjugation is used to convert a free oligonucleotide binder into a controlled nanosurface construct that can deliver stronger optical response, higher local probe density, easier separation, and more robust interaction with complex sample matrices. In practice, this helps research teams address common gaps such as weak color change in aggregation assays, unstable probes in saline or serum-containing media, poor target access caused by surface crowding, uncontrolled aptamer-to-particle ratios, and inconsistent signal transfer when moving from feasibility studies to application-focused assay formats.
A well-designed conjugate strategy considers aptamer sequence architecture, terminal handle, spacer design, nanoparticle size, surface blocking, purification route, and intended readout mechanism together rather than as isolated variables. That is especially important when the same conjugate must remain functional across binding studies, incubation steps, wash procedures, storage, and downstream integration into strips, plates, electrodes, or cell-based workflows.
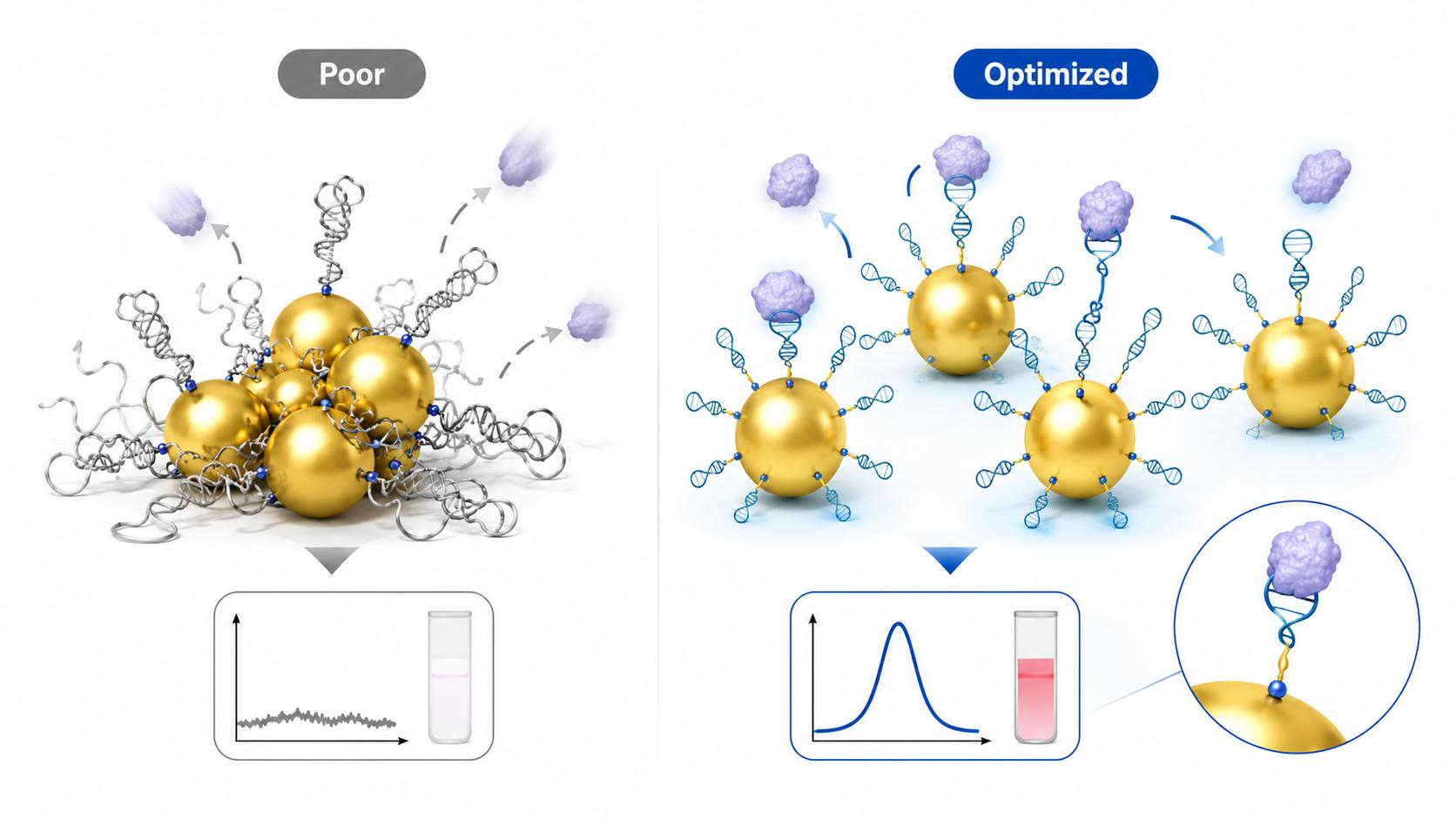 Schematic illustration of aptamer-functionalized gold nanoparticles optimized to reduce aggregation, improve target accessibility, and enhance assay signal consistency.
Schematic illustration of aptamer-functionalized gold nanoparticles optimized to reduce aggregation, improve target accessibility, and enhance assay signal consistency.Key Challenges Research Teams Face in Aptamer–AuNP Projects
Gold binding can change aptamer presentation if the terminal handle, spacer length, or grafting density is poorly matched to the sequence. We help control attachment orientation and surface spacing so the recognition region remains accessible instead of being partially flattened, sterically blocked, or forced into an unfavorable fold.
Citrate-stabilized AuNPs are sensitive to ionic strength, pH, and competing biomolecules. Projects often encounter premature aggregation during salt-aging, storage, sample incubation, or strip assembly. We optimize conjugation and passivation conditions to reduce nonspecific aggregation while preserving the response mechanism required by the intended assay.
Differences in nanoparticle size distribution, oligo reduction state, surface chemistry, and purification conditions can produce major lot-to-lot variation. We build workflows around defined particle specifications, controlled aptamer input, and loading verification to improve reproducibility across screening and follow-up batches.
A UV-Vis shift alone rarely tells you whether the conjugate is actually fit for binding or sensing. We combine physicochemical characterization with function-relevant checks—such as loading assessment, salt challenge behavior, target-binding comparison, and stability testing—so the data package supports decision-making rather than only confirming that coupling occurred.
Our Aptamer Gold Nanoparticle Conjugation Services
We provide custom service packages for aptamer-functionalized gold nanoparticles ranging from sequence-level planning to assay-format optimization. Projects may start from a customer-supplied aptamer, a known literature aptamer requiring reformatting, or an existing gold nanoparticle probe that needs better stability, loading control, or functional readout.
 Aptamer Design Review & Terminal Modification Planning
Aptamer Design Review & Terminal Modification Planning
Capabilities include:
- Review of DNA or RNA aptamer sequence architecture, target-binding region, and structural constraints
- Selection of 5′ or 3′ terminal modification strategies such as thiol, disulfide, polyA anchor, biotin, fluorophore, or orthogonal coupling handles where appropriate
- Spacer design using polyT, PEG-like, or other separation motifs to improve target accessibility on the gold surface
- Reduction and activation planning for sulfur-containing handles before conjugation
- Assessment of whether direct surface immobilization, hybridization-based display, or secondary coupling is the better route for the intended application
Typical applications:
Colorimetric aptasensors, lateral flow probe design, cell-surface recognition studies, and sequence reformatting for nanoparticle presentation
 Gold Nanoparticle Selection & Surface Preparation
Gold Nanoparticle Selection & Surface Preparation
Capabilities include:
- Selection of AuNP diameter and format based on readout mechanism, transport behavior, and required surface area
- Support for citrate-stabilized or pre-functionalized particles depending on downstream chemistry
- Evaluation of particle concentration, dispersion buffer, and preconditioning requirements prior to aptamer attachment
- Integration of passivating or blocking components to balance colloidal stability with aptamer performance
- Optional coordination with broader nanoparticles and beads conjugation workflows when comparative nanoparticle formats are being assessed
Typical applications:
Visible colorimetric probes, rapid test labels, multivalent recognition constructs, and gold nanoparticle platforms for analytical method development
 Conjugation, Passivation & Functional Payload Integration
Conjugation, Passivation & Functional Payload Integration
Capabilities include:
- Thiol–gold conjugation using controlled adsorption and salt-aging style workflows when suitable
- Alternative attachment strategies such as polyA-mediated adsorption, streptavidin-biotin assembly, or click-enabled secondary coupling for specialized constructs
- Surface backfilling and passivation to manage nonspecific adsorption and storage behavior
- Co-display or hybridization of reporters, helper strands, quenchers, or cargo oligonucleotides as required by the study design
- Optimization of aptamer surface density for a balance between binding access, signal generation, and colloidal stability
Focus areas:
Aggregation control, accessible aptamer presentation, matrix tolerance, and translation into assay-ready or uptake-study-ready conjugates
 Analytical Characterization & Assay-Fit Optimization
Analytical Characterization & Assay-Fit Optimization
Capabilities include:
- UV-Vis confirmation of plasmon profile and aggregation behavior
- DLS and zeta potential assessment to monitor hydrodynamic size and surface-charge changes after conjugation
- TEM or related morphology review where required
- Aptamer loading or aptamer-to-particle ratio estimation using appropriate quantification strategies
- Salt challenge, storage evaluation, and matrix compatibility testing
- Functional comparison of candidate conjugates in binding, response, or capture experiments relevant to the target application
Deliverables:
Conjugation summary, analytical readouts, stability observations, and recommended conditions for scale-up or downstream assay integration
Key Design Parameters for Aptamer Gold Nanoparticle Conjugation
Successful aptamer–AuNP design depends on the relationship between sequence engineering, nanoparticle properties, and the intended readout mechanism. The table below highlights the variables that most often determine whether a conjugate performs well in real workflows rather than only under ideal screening conditions.
| Design Parameter | Common Options | Development Considerations | Impact on Conjugate Performance | Why It Matters to Customers |
| Aptamer Format | DNA aptamer, RNA aptamer, truncated aptamer, structure-stabilized variant | Sequence length and folding behavior influence surface presentation and target access | Affects affinity retention, nuclease tolerance, and assay robustness | Determines whether a literature sequence can be translated into a practical nanoparticle construct |
| Attachment Handle | 5′-thiol, 3′-thiol, disulfide precursor, polyA anchor, biotin, orthogonal click handle | Must align with aptamer folding and downstream chemistry | Controls surface orientation, coupling efficiency, and long-term stability | Reduces the risk of losing binding performance after immobilization |
| Gold Nanoparticle Size & Format | Small spherical AuNPs, larger spherical AuNPs, prefunctionalized AuNP formats | Surface area, optical behavior, and transport properties vary with particle design | Influences signal intensity, loading capacity, and assay compatibility | Helps match the conjugate to colorimetric, strip-based, imaging, or cell-interaction studies |
| Spacer & Passivation Strategy | PolyT spacer, PEG-like spacer, short thiol backfiller, mixed surface blocking system | Used to separate the recognition sequence from the gold surface and limit nonspecific adsorption | Improves accessibility and stabilizes the colloid in working buffers | Often determines whether the probe survives real sample handling conditions |
| Aptamer Surface Density | Low, moderate, or high loading depending on sequence and application | Excess density can crowd the binding domain, while low density can reduce signal or capture efficiency | Balances target recognition with particle stability and multivalency | A key optimization point when moving from concept data to reproducible builds |
| Working Buffer & Storage Conditions | Low-salt preparation buffer, salt-aged system, assay buffer with blockers, short-term storage buffer | Ionic strength and competing biomolecules can shift both stability and target response | Affects aggregation risk, background signal, and shelf handling behavior | Directly impacts whether the conjugate remains usable outside a single freshly prepared batch |
Aptamer–AuNP Conjugation Strategies & Process Development Considerations
There is no single attachment route that fits every aptamer and every nanoparticle format. Method selection should be driven by sequence chemistry, desired loading density, time constraints, storage needs, and the assay principle the conjugate is expected to support.
| Conjugation Strategy | Technical Approach | Common Applications | Development Advantages |
| Direct Thiol–Gold Coupling | Thiol- or disulfide-modified aptamers are attached to AuNP surfaces through Au–S chemistry | General-purpose aptamer probes, stable nanoparticle labels, multivalent recognition constructs | Widely used, strong surface binding, suitable for controlled loading workflows |
| Salt-Aging Assisted Loading | Gradual ionic-strength increase is used to improve packing density of negatively charged oligonucleotides on citrate-stabilized AuNPs | Dense aptamer loading for colorimetric, lateral flow, and hybridization-based systems | Enables higher surface coverage, but requires careful aggregation control |
| PolyA-Mediated Adsorption | Polyadenine anchor segments are used to orient the functional aptamer block away from the gold surface | Rapid build screening, tunable display architectures, some plasmonic biosensor formats | Useful for tuning spacing and presentation without relying only on terminal thiol placement |
| Biotin–Streptavidin Assembly | Biotinylated aptamers are assembled onto streptavidin-modified gold nanoparticle surfaces | Modular probe construction and rapid interchange of multiple aptamer candidates | Flexible and convenient for screening, especially when direct gold contact is undesirable |
| Click-Enabled Secondary Coupling | Orthogonal reactive handles are introduced to connect aptamers after preliminary surface functionalization | Multifunctional probes, dual-ligand systems, reporter-integrated conjugates | Expands design flexibility when direct adsorption is not ideal for the final construct |
| Hybridization-Based Display | A capture strand is immobilized on AuNPs and the aptamer module is displayed through complementary hybridization | Switchable probes, modular sensor design, interchangeable recognition sequences | Simplifies aptamer swapping and can separate the recognition element from the gold surface |
Analytical Characterization & Quality Control Framework for Aptamer–AuNP Conjugates
For aptamer-functionalized gold nanoparticles, analytical quality is not limited to proving that the oligonucleotide is present on the particle. It must also show whether the conjugate is physically stable, sufficiently loaded, and still functionally relevant to the target-binding mechanism the project depends on.
| Analytical Category | Methodology | Purpose in Development | Data Delivered |
| Optical Profile Confirmation | UV-Vis spectroscopy | Monitoring plasmon peak position, line shape, and target- or salt-induced aggregation behavior | Absorbance spectra, peak ratio trends, conjugate comparison plots |
| Hydrodynamic Size Assessment | Dynamic light scattering (DLS) | Detecting size increase after aptamer attachment and identifying aggregation tendencies | Size distribution reports and comparative hydrodynamic diameter data |
| Surface Charge Evaluation | Zeta potential analysis | Confirming surface-state change after oligonucleotide loading and passivation | Zeta potential values and batch-to-batch comparison data |
| Morphology Review | TEM or equivalent particle imaging | Verifying particle integrity, size distribution, and gross aggregation | Representative particle images and morphology notes |
| Aptamer Loading Estimation | Supernatant depletion analysis, labeled-oligo quantification, or other suitable loading assays | Estimating aptamer-to-particle ratio and comparing loading conditions | Loading efficiency summary and calculated surface coverage trends |
| Stress & Stability Testing | Salt challenge, pH tolerance, storage observation, or matrix exposure studies | Evaluating whether the conjugate remains usable under realistic handling conditions | Stability observations, spectra shifts, and recommended operating windows |
| Functional Binding Verification | Target response testing in the relevant assay format | Confirming that conjugation did not eliminate recognition performance | Comparative binding or signal response results for candidate conjugates |
| Documentation Package | Structured reporting of build conditions, analytics, and recommended handling parameters | Supporting project transfer, repeat ordering, or downstream assay development | Conjugation record, analytical summary, and condition recommendations |
Workflow for Custom Aptamer Gold Nanoparticle Conjugation
We start by clarifying the target type, assay or study format, working matrix, desired signal mode, and whether you already have an aptamer and gold nanoparticle system in hand. This step prevents sequence chemistry and particle selection from being optimized in the wrong direction.
The aptamer is reviewed for truncation, terminal handle placement, spacer needs, and any reporter or helper-strand requirements. Where necessary, alternative architectures are proposed to improve surface display without changing the intended recognition logic.
Gold nanoparticle format, concentration, buffer, and attachment route are selected and then optimized through controlled conjugation experiments. The goal is to reach a stable loading window rather than maximizing aptamer density at the expense of usability.
Unbound aptamer and unstable species are removed, and the surface is conditioned for the intended working environment. This step is especially important for projects involving salts, proteins, strip membranes, or repeated incubations.
Candidate conjugates are compared using the most relevant analytical and functional checks, such as optical response, DLS, zeta potential, loading assessment, and target-response behavior. This allows rational selection of a build that is both measurable and useful.
Final output may include research-grade conjugates, condition recommendations, handling guidance, and analytical summaries to support assay development, method transfer, or the next optimization cycle.
Why Choose Our Aptamer–AuNP Conjugation Platform
We do not treat aptamers as generic oligos. Handle placement, spacer design, folding behavior, and target access are considered together with nanoparticle chemistry so the final construct is designed for function rather than simple attachment.

Many aptamer–AuNP systems appear successful at the bench but fail during storage, sample exposure, or assay integration. Our development logic emphasizes colloidal stability, passivation, and usable operating windows from the start.
We connect UV-Vis, DLS, zeta potential, morphology, and loading data to actual target-response questions, helping teams decide which conjugate is worth advancing instead of only documenting a surface modification event.
We can support direct aptamer-coated gold nanoparticles, modular assemblies, reporter-integrated constructs, and projects that need coordination with related gold or oligonucleotide labeling workflows instead of a one-format-only service model.
Common Research Applications of Aptamer Gold Nanoparticle Conjugates
Colorimetric Aptasensors
- Target-responsive aggregation or dispersion systems for visible readout.
- Probe design for absorbance-ratio based detection workflows.
- Optimization of response window, background stability, and salt behavior.
Lateral Flow & Rapid Test Labels
- Aptamer-functionalized AuNP probes for strip-based capture and signal generation.
- Surface conditioning to improve membrane compatibility and reduce nonspecific loss.
- Comparative build support for single-aptamer and sandwich-style formats.
Cell Recognition & Uptake Studies
- Multivalent gold nanoparticle display of cell-binding aptamers.
- Constructs for receptor-focused binding, internalization, or blocking studies.
- Integration with labeled oligos or payload strands for mechanistic experiments.
Imaging, Electrochemical & SERS Probe Development
- Gold nanoparticle conjugates adapted for plasmonic, fluorescence-coupled, electrochemical, or SERS-related readouts.
- Surface engineering to support reporter integration and signal amplification.
- Custom conjugate preparation for method development and proof-of-concept studies.
Discuss Your Aptamer Gold Nanoparticle Conjugation Project
Whether you are building a new aptamer-functionalized gold nanoparticle probe, troubleshooting aggregation in an existing construct, or preparing a conjugate for a defined biosensing or targeting workflow, we provide technically focused support across design, conjugation, and characterization.
Our team works with customer-defined sequences, particle formats, and application goals to deliver conjugates and data packages that are easier to evaluate, reproduce, and integrate into downstream research.Contact our scientific team to discuss your aptamer gold nanoparticle conjugation requirements and request a project-specific proposal.
Frequently Asked Questions (FAQ)
It is the controlled attachment of a DNA or RNA aptamer to a gold nanoparticle to create a functional probe for recognition, capture, signaling, imaging-related method development, or delivery research.
The most common route uses thiol- or disulfide-modified aptamers because sulfur has strong affinity for gold. Depending on the project, polyA adsorption, biotin–streptavidin assembly, or secondary click-style coupling may also be used.
Not always. Thiol is often preferred for direct Au surface coupling, but alternative handles may be better when modular assembly, faster screening, or indirect surface presentation is needed.
Aggregation is usually caused by a mismatch among salt ramp, particle concentration, oligo reduction state, aptamer loading density, and surface blocking conditions. It can also happen when the working buffer is not compatible with the original colloid.
Size should be selected based on the intended readout, required surface area, membrane transport needs, and stability window. A particle that works for visible color development may not be ideal for uptake or electrochemical studies.

